
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,081,020 – 5,081,136 |
| Length | 116 |
| Max. P | 0.830402 |

| Location | 5,081,020 – 5,081,136 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.66 |
| Mean single sequence MFE | -43.62 |
| Consensus MFE | -34.81 |
| Energy contribution | -34.69 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.548280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5081020 116 + 23771897 UGACUAUGUGCAGUGCUAGUGUGUGUGUG----CUGAGUCCUUGUGUGGGUGCACUGGCAUGUUGUGAUUGCGGACUGUUGCUUCACUUGAGUGGCGGCAAGUCGGCGACCCAACAGCUA .........((.(((((((((..(.((..----(.........)..)).)..)))))))))((((.(.((((.(((((((((..(((....)))))))).)))).)))).))))).)).. ( -44.70) >DroSec_CAF1 287286 116 + 1 UACCUAUGUGCAGUGCAAAUGUGUGUGUG----CUGAGUCCUUGUGUGGGUGCACUGGCAUGUUGUGAUUGCGGACUGUUGCUUCACUUGAGUGGCGACAAGUCGGCGACCAAACGGCUA ((((((..(.(((..((........))..----))).......)..))))))...((((.((((.((.((((.(((((((((..(((....)))))))).)))).)))).)))))))))) ( -38.10) >DroSim_CAF1 308630 116 + 1 UGCCUAUGUGCAGUGCAAAUGUGUGUGUG----CUGAGUCCUUGUGUGGGUGCACUGGCAUGUUGUGAUUGCGGACUGUUGCUUCACUUGAGUGGCGGCAAGUCGGCGACCCAACGGCUA .(((.((((.(((((((...((......)----)....(((......))))))))))))))((((.(.((((.(((((((((..(((....)))))))).)))).)))).)))))))).. ( -42.40) >DroYak_CAF1 298724 120 + 1 UGCCUAUGUACAGUGCCAGUGUGUGUGUGUGAGUUGAGUCCUUGUGUGGGUGCACUGGCAUGUUGUGAUUGCGGACUGUUGCUUCACUUCAGUGGCAGCAAGUCGGCGACCCAACAGCAA (((.........(((((((((..(.((..((((.......))))..)).)..)))))))))((((.(.((((.(((((((((..(((....)))))))).)))).)))).))))).))). ( -49.30) >consensus UGCCUAUGUGCAGUGCAAAUGUGUGUGUG____CUGAGUCCUUGUGUGGGUGCACUGGCAUGUUGUGAUUGCGGACUGUUGCUUCACUUGAGUGGCGGCAAGUCGGCGACCCAACAGCUA .....((((.(((((((.....................(((......))))))))))))))(((((..((((.(((((((((..(((....)))))))).)))).))))....))))).. (-34.81 = -34.69 + -0.12)



| Location | 5,081,020 – 5,081,136 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.66 |
| Mean single sequence MFE | -32.24 |
| Consensus MFE | -26.19 |
| Energy contribution | -25.88 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830402 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5081020 116 - 23771897 UAGCUGUUGGGUCGCCGACUUGCCGCCACUCAAGUGAAGCAACAGUCCGCAAUCACAACAUGCCAGUGCACCCACACAAGGACUCAG----CACACACACACUAGCACUGCACAUAGUCA ..((((((((((.((.(((((((...(((....)))..)))..)))).)).))).))))).))((((((..((......)).....(----......)......)))))).......... ( -29.10) >DroSec_CAF1 287286 116 - 1 UAGCCGUUUGGUCGCCGACUUGUCGCCACUCAAGUGAAGCAACAGUCCGCAAUCACAACAUGCCAGUGCACCCACACAAGGACUCAG----CACACACACAUUUGCACUGCACAUAGGUA ..((((((((((.((.((((.((.(((((....)))..)).)))))).)).)))).))).(((.((((((.((......)).....(----......).....)))))))))....))). ( -29.80) >DroSim_CAF1 308630 116 - 1 UAGCCGUUGGGUCGCCGACUUGCCGCCACUCAAGUGAAGCAACAGUCCGCAAUCACAACAUGCCAGUGCACCCACACAAGGACUCAG----CACACACACAUUUGCACUGCACAUAGGCA ..((((((((((.((.(((((((...(((....)))..)))..)))).)).))).)))).(((.((((((.((......)).....(----......).....)))))))))....))). ( -33.90) >DroYak_CAF1 298724 120 - 1 UUGCUGUUGGGUCGCCGACUUGCUGCCACUGAAGUGAAGCAACAGUCCGCAAUCACAACAUGCCAGUGCACCCACACAAGGACUCAACUCACACACACACACUGGCACUGUACAUAGGCA .(((((((((((.((.((((((((..(((....))).))))..)))).)).))).)))).((((((((..........((.......))..........)))))))).........)))) ( -36.15) >consensus UAGCCGUUGGGUCGCCGACUUGCCGCCACUCAAGUGAAGCAACAGUCCGCAAUCACAACAUGCCAGUGCACCCACACAAGGACUCAG____CACACACACACUUGCACUGCACAUAGGCA ..((((((((((.((.(((((((...(((....)))..)))..)))).)).))).)))).(((.(((((...................................))))))))....))). (-26.19 = -25.88 + -0.31)

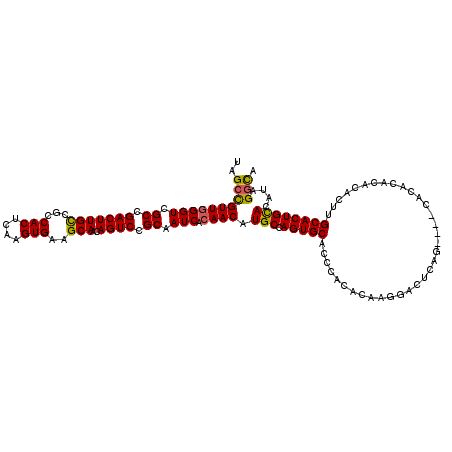

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:00 2006