
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,071,528 – 5,071,682 |
| Length | 154 |
| Max. P | 0.990050 |

| Location | 5,071,528 – 5,071,647 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.40 |
| Mean single sequence MFE | -24.79 |
| Consensus MFE | -21.00 |
| Energy contribution | -20.92 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955171 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5071528 119 + 23771897 AGCAAACCAAACCAGUGGGAA-AAAAUUGUUGUUGACGGCAAGAAUUUUCCACAUUCAUACACUGAACCAGCAAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCUUGUGAC .((...........(((((((-(...((((((....))))))...)))))))).((((.....))))...)).((((.(((((....))))).......((........))))))..... ( -25.70) >DroSec_CAF1 278046 119 + 1 AGCAAACCAAACCAGUGGGAA-AAAAUUGUUGUUGACGGCAAGAAUUUUCCACAUUCAUACACUGAACCGGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGAC ..............(((((((-(...((((((....))))))...))))))))..((((((.((......((((....(((((....))))).....)))).......)).)..))))). ( -24.92) >DroSim_CAF1 299327 119 + 1 AGCAAACCAAACCAGUGGGAA-AAAAUUGUUGUUGACGGCAAGAAUUUUCCACAUUCAUACACUGAACCGGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGAC ..............(((((((-(...((((((....))))))...))))))))..((((((.((......((((....(((((....))))).....)))).......)).)..))))). ( -24.92) >DroEre_CAF1 284965 120 + 1 AGAAAACCAAACCAGUGCGAAAAAAAUUGUUGUUGACGGCAAGAAUUUUUCACAUUCAUACAGUGAGCCAGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCUUGUGAC ..........((.(((((((((((..((((((....))))))...))))))...........(((((...((...)).(((((....)))))....)))))........))))).))... ( -25.20) >DroYak_CAF1 288865 119 + 1 AGCAAACCAAACCAGUGGGAA-AAAAUUGUUGUUGACGGCAAGAAUUUUCCACAUUCAUACACUGAACCAGCCAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGAC .((...........(((((((-(...((((((....))))))...)))))))).((((.....))))...))((.((.(((((....))))).......((........)))).)).... ( -23.20) >consensus AGCAAACCAAACCAGUGGGAA_AAAAUUGUUGUUGACGGCAAGAAUUUUCCACAUUCAUACACUGAACCAGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGAC ..............(((((((.....((((((....))))))....))))))).............((.(((......(((((....))))).......((........))))).))... (-21.00 = -20.92 + -0.08)


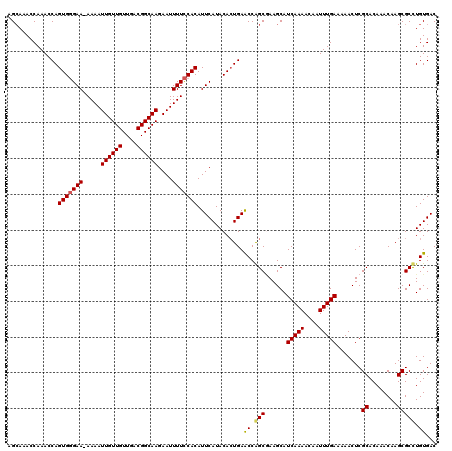
| Location | 5,071,567 – 5,071,682 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.76 |
| Mean single sequence MFE | -20.62 |
| Consensus MFE | -16.50 |
| Energy contribution | -16.34 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911346 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5071567 115 + 23771897 AAGAAUUUUCCACAUUCAUACACUGAACCAGCAAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCUUGUGACAGGCAAAAAUGGAAAUGCAAUAAAAUGA-----AUGGCAA ............(((((((...(((.((.(((......(((((....))))).......((........))))).))..)))(((..........)))......))))-----))).... ( -19.50) >DroSec_CAF1 278085 115 + 1 AAGAAUUUUCCACAUUCAUACACUGAACCGGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGACAGGCAAAAAUGGAAAUGCAAUAAAAUGA-----AUGGCGA ............(((((((...........((((....(((((....))))).....))))........(((((((...)))))....((....))))......))))-----))).... ( -22.50) >DroSim_CAF1 299366 115 + 1 AAGAAUUUUCCACAUUCAUACACUGAACCGGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGACAGGCAAAAAUGGAAAUGCAAUAAAAUGA-----AUGGCGA ............(((((((...........((((....(((((....))))).....))))........(((((((...)))))....((....))))......))))-----))).... ( -22.50) >DroEre_CAF1 285005 115 + 1 AAGAAUUUUUCACAUUCAUACAGUGAGCCAGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCUUGUGACAGGCAAAAAUGGAAAUGCAAUAAAAUGA-----AUGGCGA ............(((((((....((..(((((((....(((((....))))).....))))..........(((((...))))).....))).....)).....))))-----))).... ( -21.60) >DroYak_CAF1 288904 120 + 1 AAGAAUUUUCCACAUUCAUACACUGAACCAGCCAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGACAGGCACAAAUGGAAAUGCAAUAAAAUUAAUAAAAUGAAAA ..............(((((...(((...)))....((((.......(((((........((........))(((((...))))).)))))....))))...............))))).. ( -17.00) >consensus AAGAAUUUUCCACAUUCAUACACUGAACCAGCGAAGCAUCAAAACAAUUUGAAAAACUCGCACAAACAAGCGCCUGUGACAGGCAAAAAUGGAAAUGCAAUAAAAUGA_____AUGGCGA ..(...((((((..((((.....))))...((((....(((((....))))).....))))..........(((((...))))).....))))))..)...................... (-16.50 = -16.34 + -0.16)

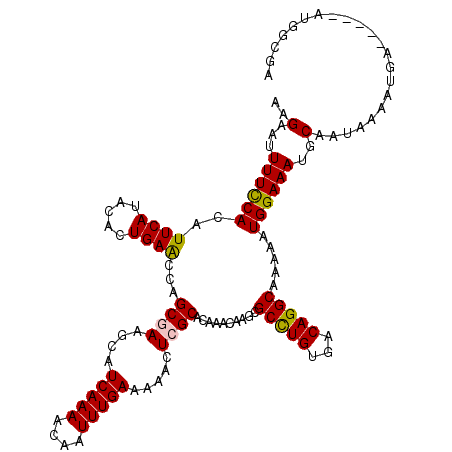

| Location | 5,071,567 – 5,071,682 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.76 |
| Mean single sequence MFE | -27.10 |
| Consensus MFE | -23.24 |
| Energy contribution | -22.80 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.990050 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5071567 115 - 23771897 UUGCCAU-----UCAUUUUAUUGCAUUUCCAUUUUUGCCUGUCACAAGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUUGCUGGUUCAGUGUAUGAAUGUGGAAAAUUCUU .(((.((-----.......)).)))(((((((...((((((...((.((((......))))((((...(((((....))))).))))...))...))).))).....)))))))...... ( -24.60) >DroSec_CAF1 278085 115 - 1 UCGCCAU-----UCAUUUUAUUGCAUUUCCAUUUUUGCCUGUCACAGGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUCGCCGGUUCAGUGUAUGAAUGUGGAAAAUUCUU ((..(((-----((((..(((((.....((......(((((...)))))........(.((((((...(((((....))))).)).)))))))..))))).)))))))..))........ ( -28.20) >DroSim_CAF1 299366 115 - 1 UCGCCAU-----UCAUUUUAUUGCAUUUCCAUUUUUGCCUGUCACAGGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUCGCCGGUUCAGUGUAUGAAUGUGGAAAAUUCUU ((..(((-----((((..(((((.....((......(((((...)))))........(.((((((...(((((....))))).)).)))))))..))))).)))))))..))........ ( -28.20) >DroEre_CAF1 285005 115 - 1 UCGCCAU-----UCAUUUUAUUGCAUUUCCAUUUUUGCCUGUCACAAGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUCGCUGGCUCACUGUAUGAAUGUGAAAAAUUCUU ((((.((-----((((......(((..........)))..(((((((((....))))))((((((...(((((....))))).)).)))).))).......))))))))))......... ( -24.50) >DroYak_CAF1 288904 120 - 1 UUUUCAUUUUAUUAAUUUUAUUGCAUUUCCAUUUGUGCCUGUCACAGGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUGGCUGGUUCAGUGUAUGAAUGUGGAAAAUUCUU .........................(((((((..(((((((...)))))))..(((..(((((((...(((((....))))).)))).((((...)))))))..))))))))))...... ( -30.00) >consensus UCGCCAU_____UCAUUUUAUUGCAUUUCCAUUUUUGCCUGUCACAGGCGCUUGUUUGUGCGAGUUUUUCAAAUUGUUUUGAUGCUUCGCUGGUUCAGUGUAUGAAUGUGGAAAAUUCUU .........................((((((((..((((((.....(((((((((....))))))...(((((....)))))......)))....))).)))..)))).))))....... (-23.24 = -22.80 + -0.44)


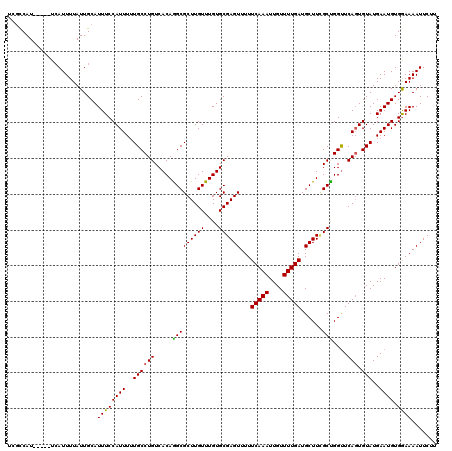
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:35 2006