
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,042,870 – 5,042,970 |
| Length | 100 |
| Max. P | 0.629295 |

| Location | 5,042,870 – 5,042,970 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 96.30 |
| Mean single sequence MFE | -22.58 |
| Consensus MFE | -18.58 |
| Energy contribution | -19.46 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.629295 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
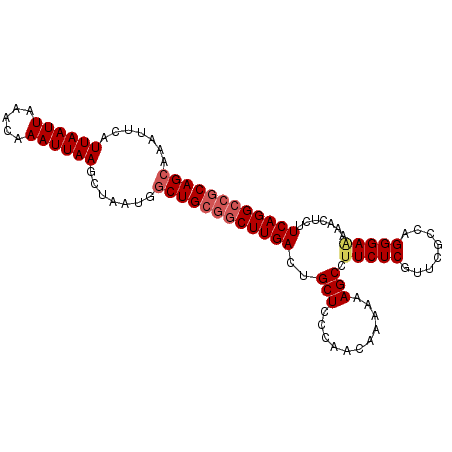
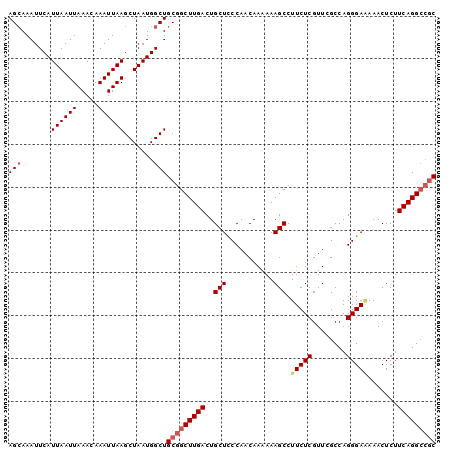
>3L_DroMel_CAF1 5042870 100 + 23771897 AGAAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUCCCAACAAAAAAGCCUUCUCGUUCGCCAGGGAAAAACUCUUCAGGCCGC .......((((((.((((.....))))..))))))..(((((((((..(((...........))).(((((........))))).......))))))))) ( -21.90) >DroSec_CAF1 249344 100 + 1 AGCAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUCCCAACAAAAAAGCCUUCUCGUUCGCCAGGGAAAAACUCUUCAGGCCGC (((.......((((((.....)))))).......)))(((((((((..(((...........))).(((((........))))).......))))))))) ( -23.44) >DroSim_CAF1 270213 97 + 1 AGCAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUCCCAACAAAAAAGCCUUCUCGUUCGCCAGGGAAAAACUCUUCAGG---C .(((...((((((.((((.....))))..)))))).)))(((((...((.......))...))))).........(((((((.....))))...))---) ( -16.60) >DroEre_CAF1 252790 99 + 1 AGCAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUCCCAACAAGAAAGCCUUCUCGUUUGCCAGGGA-AAACUCUUCAGGCCGC (((.......((((((.....)))))).......)))(((((((((....(((((((.((((....)))).))).....))))-.......))))))))) ( -25.34) >DroYak_CAF1 259193 100 + 1 AGCAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUUCCAACAAAAAAGCCUUCUCGUUUGCCAGGGAGAAACUCUUCAGGCCGC (((.......((((((.....)))))).......)))(((((((((...(((((..((((..((....))..))))...))))).......))))))))) ( -25.64) >consensus AGCAAAUUCAUUAAUUAAACAAAUUAAGCUAAUGGCUGCGGCUUGACUGCUCCCAACAAAAAAGCCUUCUCGUUCGCCAGGGAAAAACUCUUCAGGCCGC (((.......((((((.....)))))).......)))(((((((((..(((...........))).(((((........))))).......))))))))) (-18.58 = -19.46 + 0.88)

| Location | 5,042,870 – 5,042,970 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 96.30 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -24.52 |
| Energy contribution | -25.28 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.21 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.556190 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
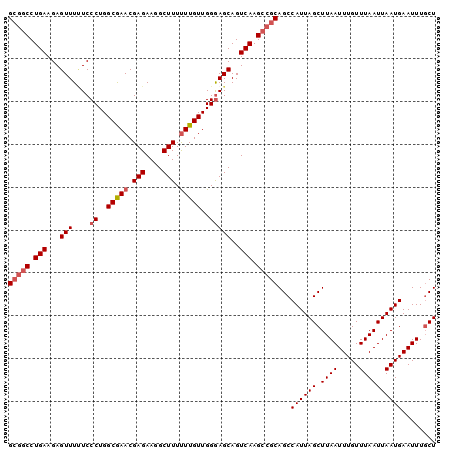
>3L_DroMel_CAF1 5042870 100 - 23771897 GCGGCCUGAAGAGUUUUUCCCUGGCGAACGAGAAGGCUUUUUUGUUGGGAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUUCU (((((.(((...(((....((..(((((.(((....))).)))))..)))))..))).)))))...((((((.((((.....))))))))))........ ( -28.60) >DroSec_CAF1 249344 100 - 1 GCGGCCUGAAGAGUUUUUCCCUGGCGAACGAGAAGGCUUUUUUGUUGGGAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUGCU (((((.(((...(((....((..(((((.(((....))).)))))..)))))..))).))))).......(((..((((..((.....))..)))).))) ( -29.70) >DroSim_CAF1 270213 97 - 1 G---CCUGAAGAGUUUUUCCCUGGCGAACGAGAAGGCUUUUUUGUUGGGAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUGCU (---((.((((....))))...))).........(((((..((((.....))))..)))))((((.((((((.((((.....))))))))))...)))). ( -21.50) >DroEre_CAF1 252790 99 - 1 GCGGCCUGAAGAGUUU-UCCCUGGCAAACGAGAAGGCUUUCUUGUUGGGAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUGCU (((((.(((...((..-((((.....((((((((....))))))))))))))..))).))))).......(((..((((..((.....))..)))).))) ( -29.00) >DroYak_CAF1 259193 100 - 1 GCGGCCUGAAGAGUUUCUCCCUGGCAAACGAGAAGGCUUUUUUGUUGGAAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUGCU (((((.(((...((((((....((((((.(((....))).))))))))))))..))).))))).......(((..((((..((.....))..)))).))) ( -28.60) >consensus GCGGCCUGAAGAGUUUUUCCCUGGCGAACGAGAAGGCUUUUUUGUUGGGAGCAGUCAAGCCGCAGCCAUUAGCUUAAUUUGUUUAAUUAAUGAAUUUGCU (((((.(((...(((....((..(((((.(((....))).)))))..)))))..))).)))))...((((((.((((.....))))))))))........ (-24.52 = -25.28 + 0.76)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:15:15 2006