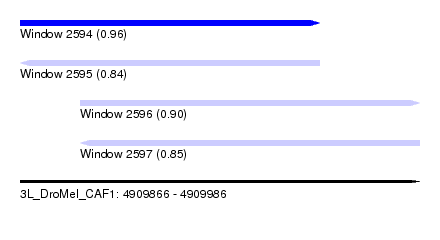
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,909,866 – 4,909,986 |
| Length | 120 |
| Max. P | 0.963320 |

| Location | 4,909,866 – 4,909,956 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 77.97 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -21.91 |
| Energy contribution | -23.80 |
| Covariance contribution | 1.89 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963320 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4909866 90 + 23771897 AUGGGUGG-----------CU-----------CAAAAGUGGAAUGGCCAGCUG-------GUUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ..(..(.(-----------((-----------(((.(((((.....))).)).-------.)))))).)..)(.((((((((((......))))))))))).....((((....)))). ( -33.10) >DroVir_CAF1 168245 79 + 1 AAGGGGGG----------------------------GGGGGGGUGGGCAGCAC-------AGCGGGCGG-----GCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAACGAG .....(..----------------------------(..(.((..(((.((..-------.....)).(-----((((((((((......))))))))))))))..)).)..)..)... ( -29.40) >DroSec_CAF1 131114 90 + 1 AUGGGUGG-----------UU-----------CAGAUGUGGAAUGGCCAGCUG-------GUUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ...(((.(-----------((-----------(.......)))).))).((((-------.((((((((..((.((((((((((......)))))))))))).))))).)))))))... ( -33.80) >DroSim_CAF1 141122 90 + 1 AUGGGUGG-----------UU-----------CAGAUGUGGAAUGGCCAGCUG-------GUUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ...(((.(-----------((-----------(.......)))).))).((((-------.((((((((..((.((((((((((......)))))))))))).))))).)))))))... ( -33.80) >DroEre_CAF1 132443 93 + 1 AUGG---------------CU-----------CAGAAGUGGGAUGGCCAGCUGUUGGGUUGUUGGGCGGCUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ..((---------------((-----------(((.(((.((....)).))).)))))))((((.(((((.((.((((((((((......))))))))))))...)))))..))))... ( -37.10) >DroYak_CAF1 135255 114 + 1 AUGGGUGGUGCAAAAGUGGUGCAGAAGUGGUGCAAAAGUGGGUUGG-CAGCUGC----UUGUUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ..(((((((((....(((.(((....))).)))((((((.(.((((-..(((((----(.....)))))))))).).))))))............).)))))))).((((....)))). ( -35.60) >consensus AUGGGUGG___________UU___________CAGAAGUGGAAUGGCCAGCUG_______GUUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAG ..(((((................................(((.((.(((((.........))))).)).)))..((((((((((......))))))))))))))).((((....)))). (-21.91 = -23.80 + 1.89)


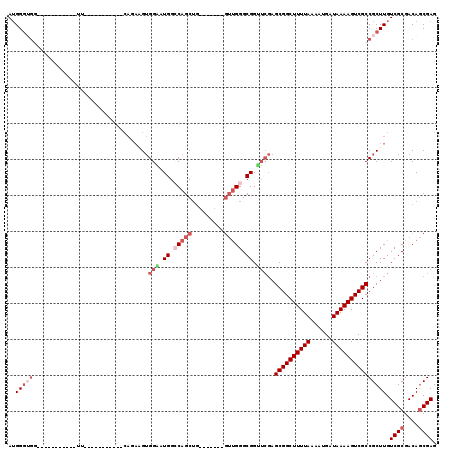
| Location | 4,909,866 – 4,909,956 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 77.97 |
| Mean single sequence MFE | -20.38 |
| Consensus MFE | -15.99 |
| Energy contribution | -16.18 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.842422 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4909866 90 - 23771897 CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAAC-------CAGCUGGCCAUUCCACUUUUG-----------AG-----------CCACCCAU ...((((..(((.....(((((.((((((......)))))).))).))...))).....-------))))((((((.........))-----------.)-----------)))..... ( -23.30) >DroVir_CAF1 168245 79 - 1 CUCGUUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGC-----CCGCCCGCU-------GUGCUGCCCACCCCCCC----------------------------CCCCCCUU ...((.(.((((.(..((((((.((((((......)))))).)).-----))))..).)-------)))).))..........----------------------------........ ( -17.30) >DroSec_CAF1 131114 90 - 1 CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAAC-------CAGCUGGCCAUUCCACAUCUG-----------AA-----------CCACCCAU ...((((..(((.....(((((.((((((......)))))).))).))...))).....-------))))(((.....)))......-----------..-----------........ ( -18.50) >DroSim_CAF1 141122 90 - 1 CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAAC-------CAGCUGGCCAUUCCACAUCUG-----------AA-----------CCACCCAU ...((((..(((.....(((((.((((((......)))))).))).))...))).....-------))))(((.....)))......-----------..-----------........ ( -18.50) >DroEre_CAF1 132443 93 - 1 CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAGCCGCCCAACAACCCAACAGCUGGCCAUCCCACUUCUG-----------AG---------------CCAU ...(((((.((.....))((((.((((((......))))))..((....))))))..........)))))((((((.........))-----------.)---------------))). ( -23.40) >DroYak_CAF1 135255 114 - 1 CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAACAA----GCAGCUG-CCAACCCACUUUUGCACCACUUCUGCACCACUUUUGCACCACCCAU ...((((((((.....)))(((.((((((......)))))).)))................----)))))((-(.((......)).))).......((((.......))))........ ( -21.30) >consensus CUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAAC_______CAGCUGGCCAUCCCACUUCUG___________AA___________CCACCCAU ...((((.(((.....)))(((.((((((......)))))).))).....................))))................................................. (-15.99 = -16.18 + 0.19)


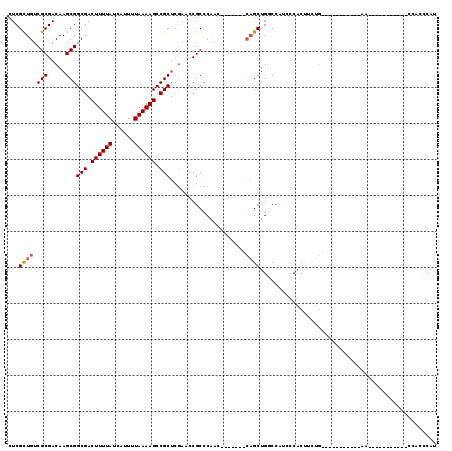
| Location | 4,909,884 – 4,909,986 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 78.96 |
| Mean single sequence MFE | -35.33 |
| Consensus MFE | -22.42 |
| Energy contribution | -23.58 |
| Covariance contribution | 1.17 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898261 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4909884 102 + 23771897 GAAUGGCCAGCUG-------G-UUGGGCGGUUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAGACGCCGACAA-AGCGGCGUAAAUUUCAAUUA ((((.(((.....-------.-...))).))))(.((((((((((......))))))))))).....((((....)))).((((((....-..))))))............ ( -36.90) >DroVir_CAF1 168257 97 + 1 GGGUGGGCAGCAC-------A-GCGGGCGG-----GCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAACGAGUCGCCGACAA-AGCGGCGUAAAUUUCAAUUA ...((.((.((..-------.-..((((((-----.(((((((((......)))))))))))))))((((((((.....)))).))))..-.)).)).))........... ( -36.60) >DroGri_CAF1 167853 90 + 1 GUAG--------C-------A-GCGGGCGG-----GCGCGCUUUUCAAUGAUAAAAGUCGCCGCAUGUCGCGACAGCGAGACGCCGACAAAAGCGGCGUAAAUUUCAAUUA ((.(--------(-------.-(((.((((-----.((.((((((.......)))))))))))).))).)).))......((((((.(....)))))))............ ( -27.90) >DroEre_CAF1 132457 109 + 1 GGAUGGCCAGCUGUUGGGUUG-UUGGGCGGCUCGAGCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAGACGCCGACAA-AACGGCGUAAAUUUCAAUUA ((....))....(((((((((-(((.(((((.((.((((((((((......))))))))))))...)))))..)))))).((((((....-..))))))....)))))).. ( -40.70) >DroMoj_CAF1 177813 93 + 1 GGACGAGCGG------------UCGGGCGG-----UCGGCUUUUAAAAUGAUAAAAGUCGAGGCUUGUCGCGACAACGAGACGCCGACAA-AGCGGCGUAAAUUUCAAUUA ....((((((------------.(((((..-----((((((((((......)))))))))).))))))))).........((((((....-..)))))).....))..... ( -32.60) >DroAna_CAF1 139335 109 + 1 GGGCGGCCUGGUGGUUGGAUGUUUGGGUGGU-UGGGCGGCUUUUAAAAUGAUAAAAGUCCCCGCUUGUCGCGACAGGGAGACGCCGACAA-AGCGGCGUAAAUUUCAAUUA ...(((((....)))))..((((..((..(.-((((.((((((((......)))))))))))).)..))..))))(..(.((((((....-..))))))...)..)..... ( -37.30) >consensus GGAUGGCCAGCUG_______G_UCGGGCGG_____GCGGCUUUUAAAAUGAUAAAAGUCGCCGCUUGUCGCGACAGCGAGACGCCGACAA_AGCGGCGUAAAUUUCAAUUA .......................((((((......((((((((((......))))))))))))))))((((....)))).((((((.......))))))............ (-22.42 = -23.58 + 1.17)


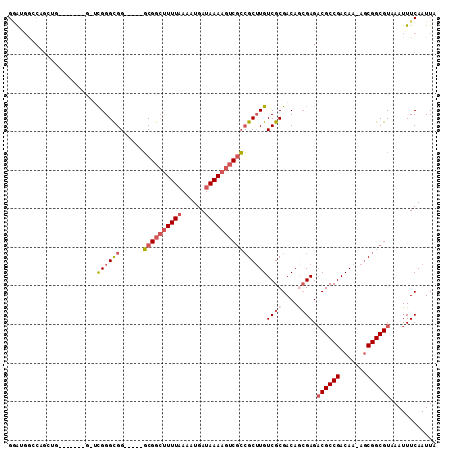
| Location | 4,909,884 – 4,909,986 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 78.96 |
| Mean single sequence MFE | -28.25 |
| Consensus MFE | -20.53 |
| Energy contribution | -21.25 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4909884 102 - 23771897 UAAUUGAAAUUUACGCCGCU-UUGUCGGCGUCUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAACCGCCCAA-C-------CAGCUGGCCAUUC ...((((......(((((((-(.((((.((.(.....).)))))))))))))).((((((......))))))....))))...(((...-.-------.....)))..... ( -30.30) >DroVir_CAF1 168257 97 - 1 UAAUUGAAAUUUACGCCGCU-UUGUCGGCGACUCGUUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGC-----CCGCCCGC-U-------GUGCUGCCCACCC .............(((((((-(.((((.((((.....)))))))))))))))).((((((......))))))..((-----.(((....-.-------)))..))...... ( -31.90) >DroGri_CAF1 167853 90 - 1 UAAUUGAAAUUUACGCCGCUUUUGUCGGCGUCUCGCUGUCGCGACAUGCGGCGACUUUUAUCAUUGAAAAGCGCGC-----CCGCCCGC-U-------G--------CUAC .............((((((...(((((.((.(.....).))))))).))))))................((((((.-----.....)))-.-------)--------)).. ( -28.40) >DroEre_CAF1 132457 109 - 1 UAAUUGAAAUUUACGCCGUU-UUGUCGGCGUCUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGCUCGAGCCGCCCAA-CAACCCAACAGCUGGCCAUCC ..............((((((-((((.((((.((((....(((.....)))(((.((((((......)))))).))).)))).))))..)-)))......))).)))..... ( -30.00) >DroMoj_CAF1 177813 93 - 1 UAAUUGAAAUUUACGCCGCU-UUGUCGGCGUCUCGUUGUCGCGACAAGCCUCGACUUUUAUCAUUUUAAAAGCCGA-----CCGCCCGA------------CCGCUCGUCC ............(((..((.-..(((((.(((.((....)).)))..((.(((.((((((......)))))).)))-----..))))))------------).)).))).. ( -21.50) >DroAna_CAF1 139335 109 - 1 UAAUUGAAAUUUACGCCGCU-UUGUCGGCGUCUCCCUGUCGCGACAAGCGGGGACUUUUAUCAUUUUAAAAGCCGCCCA-ACCACCCAAACAUCCAACCACCAGGCCGCCC ..............((((((-(.((((.((.(.....).))))))))))((((.((((((......)))))).).))).-.......................)))..... ( -27.40) >consensus UAAUUGAAAUUUACGCCGCU_UUGUCGGCGUCUCGCUGUCGCGACAAGCGGCGACUUUUAUCAUUUUAAAAGCCGC_____CCGCCCAA_C_______CAGCUGGCCAUCC .............(((((((..(((((.((.(.....).)))))))))))))).((((((......))))))....................................... (-20.53 = -21.25 + 0.72)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:35 2006