
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,758,771 – 4,758,885 |
| Length | 114 |
| Max. P | 0.650581 |

| Location | 4,758,771 – 4,758,885 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.00 |
| Mean single sequence MFE | -34.59 |
| Consensus MFE | -25.56 |
| Energy contribution | -25.37 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.650581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4758771 114 - 23771897 GAGACCCUCUGUUUGUCCAGGAUAAUGCCCGGGCGG----GAUAGUUGUUA--CCGAGUACAUAACUUAAAGCCGAAAGCAGCUCCAACACUAUUCUGCGUUAAAUUCCAGGAUAAACAG ........((((((((((.(((........(.((((----((((((.(((.--..((((.....))))..((((....)..)))..))))))))))))).).....))).)))))))))) ( -33.92) >DroSec_CAF1 21483 114 - 1 GAGACCCUCUGUUUGUCCAGGAUAAUGCCCGCGCGG----UAUAGUUGUUA--CCGAGUACAUAACUUAAAGCCGAAAGCAGCUCCUACACUAUUCUGCGUUAAAUUUCAGGACAAACAG ........((((((((((.((.......))((((((----.(((((.((..--..((((.....))))..((((....)..)))...))))))).)))))).........)))))))))) ( -32.60) >DroSim_CAF1 18817 114 - 1 GAGAACCUCUGUUUGUCCAGGAUAAUGCCCGCGCGG----GAUAGUUGUUA--CCGAGUACAUAACUUAAAGCCGAAAGCAGCUCCUACACUAUUCUGCGUUAAAUUCCAGGACAAACAG ........((((((((((.(((........((((((----((((((.((..--..((((.....))))..((((....)..)))...)))))))))))))).....))).)))))))))) ( -38.12) >DroEre_CAF1 10742 114 - 1 GAGGACCUCUGUUUGUCCAGGAUAAUGCCCGCGCGG----UGUAGUUGUUA--CCGAGUACAUAACUUAAAGCCAAAAGCAGCUCCUACACUAUUCUGCGUUAAAUUCCAGGACAAACAG ........((((((((((.(((((((((......((----(((((..(((.--..((((.....))))...((.....)))))..))))))).....))))))...))).)))))))))) ( -35.90) >DroYak_CAF1 17522 114 - 1 GAGACCCUCUGUUUGUCCAGGAUAAUGCCCGUGCAG----GAUAGUUGUUA--CCGAGUACAUAACUUAAAGCCAAAAGCAGCUCCUACACUAUUCUGCGUUAAAUUCCAGGACAAACAG ........((((((((((.(((.........(((((----((((((.((..--..((((......................))))..)))))))))))))......))).)))))))))) ( -33.91) >DroAna_CAF1 11475 110 - 1 -------AAUGUUUGCAUUGGAUGAUACCAGUGCUGGGGAGAUAGUGGUUGCGCCAAGUACAUAACUUAAAGCCAAAUGCU--UCUGGCCCCAUUC-GCGUUAAACUCCCUGGCAAACAG -------..(((((..(((((......)))))((..(((((...(((..((.(((((((.....)))).((((.....)))--)..)))..))..)-))......)))))..))))))). ( -33.10) >consensus GAGACCCUCUGUUUGUCCAGGAUAAUGCCCGCGCGG____GAUAGUUGUUA__CCGAGUACAUAACUUAAAGCCAAAAGCAGCUCCUACACUAUUCUGCGUUAAAUUCCAGGACAAACAG ........((((((((((.(((........((((((....(((((.(((......((((.....))))...((.....)).......)))))))))))))).....))).)))))))))) (-25.56 = -25.37 + -0.19)

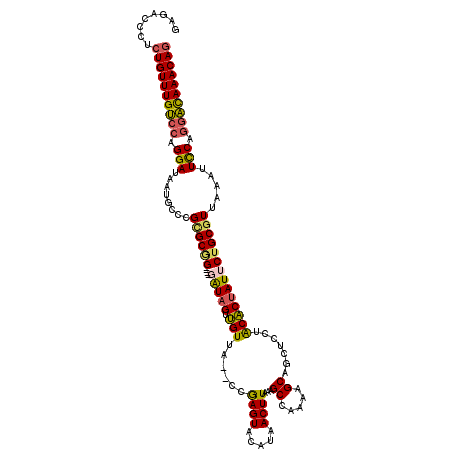
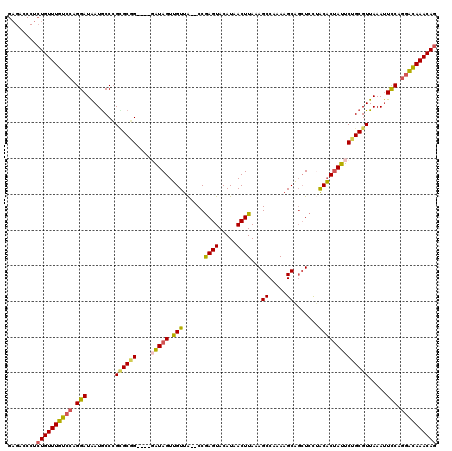
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:29 2006