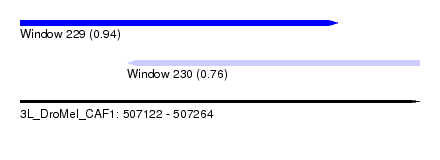

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 507,122 – 507,264 |
| Length | 142 |
| Max. P | 0.939167 |

| Location | 507,122 – 507,235 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 89.86 |
| Mean single sequence MFE | -31.88 |
| Consensus MFE | -24.42 |
| Energy contribution | -25.46 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939167 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
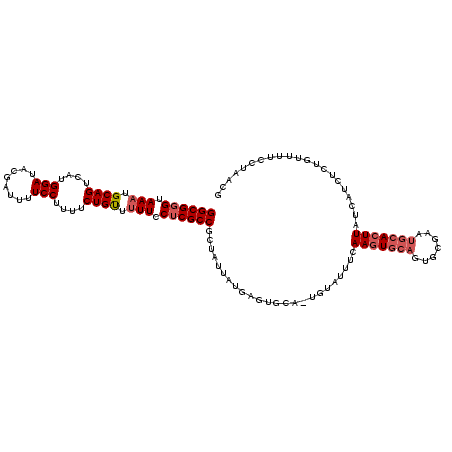
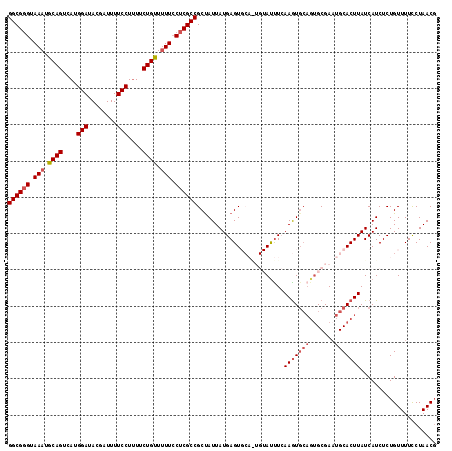
>3L_DroMel_CAF1 507122 113 + 23771897 GGCGGGUAAAUGCAGUCAUGGAUACGAUUUUCCUUUUCUGUUUUUCCUCGCCGCUAUUAUGAGUGCAGUGUAUUUCAAGU-----GCGAAUGCACUUAUCAUCUCUGUUUUCCUAACG ((((((.(((.((((....(((........)))....)))).))).))))))......(((((((((.(((((.....))-----)))..)))))))))................... ( -33.60) >DroSec_CAF1 87312 117 + 1 GGCGGGUAAAUGCAGUCAUGGAUACGAUUUUCCUUUUCUGUU-UUCCUCGCCGCCAUUAUGAGUGCAGUGUAUUUCAAGUGCAGUGCGAUUGCACUUAUCAUCUCUGUUUUCCUAACU ((((((.(((.((((....(((........)))....)))))-)).))))))......((((((((((((((((.(....).))))).)))))))))))................... ( -34.90) >DroSim_CAF1 88744 118 + 1 GGCGGGUAAAUGCAGUCAUGGAUACGAUUUUCCUUUUCUGUUUUUCCUCGCCGCUAUUAUGAGUGCAAUGUAUUUCAAGUGCAGUGCGAAUGCACUUAUCAUCUCUGUUUUCCUAACG ((((((.(((.((((....(((........)))....)))).))).))))))......(((((((((.((((((.(....).))))))..)))))))))................... ( -33.60) >DroEre_CAF1 90346 110 + 1 GGCGGGUAAAUGCAGUCAUGGAUACGAUUUUCCUUUUCUGCUUUU-CUCGCCGCUAUUAUGAGU-------ACUUGAAAUGCAGUGCGAAUGCACUUAUCAUCUCUGUUUUUCUAACG ((((((.(((.((((....(((........)))....)))).)))-))))))........(((.-------((..((.(((.(((((....)))))...))).)).))..)))..... ( -29.90) >DroYak_CAF1 92481 111 + 1 GGCGCGUAAAUGCAGUCAUGGAUACGAUUUUCCUCUUCUGUUUUUCCUCGCCGCUAUUAUGAGU-------AUCUAAAGUGCAGUGCGAAUGCACUUAUCAUCUCUGUUUUCCUAACG ((((.(.(((.((((....(((........)))....)))).))).).))))........(((.-------.....(((((((.......))))))).....)))............. ( -27.40) >consensus GGCGGGUAAAUGCAGUCAUGGAUACGAUUUUCCUUUUCUGUUUUUCCUCGCCGCUAUUAUGAGUGCA_UGUAUUUCAAGUGCAGUGCGAAUGCACUUAUCAUCUCUGUUUUCCUAACG ((((((.(((.((((....(((........)))....)))).))).))))))........................(((((((.......)))))))..................... (-24.42 = -25.46 + 1.04)

| Location | 507,160 – 507,264 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 82.11 |
| Mean single sequence MFE | -27.68 |
| Consensus MFE | -15.44 |
| Energy contribution | -16.32 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.759852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
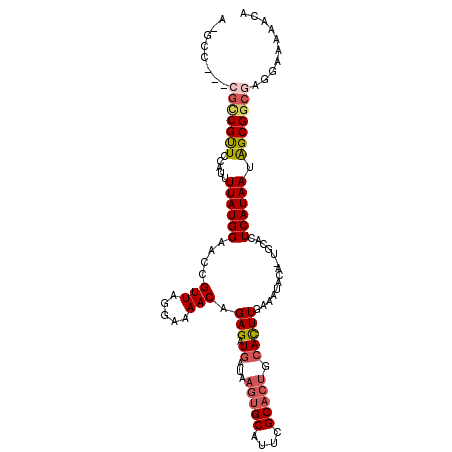
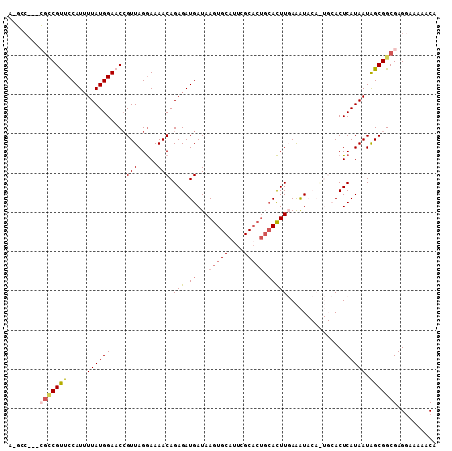
>3L_DroMel_CAF1 507160 104 - 23771897 AAGCCCGUCGUCGCUCCAUUUUAUGGAACCGUUAGGAAAACAGAGAUGAUAAGUGCAUUCGC-----ACUUGAAAUACACUGCACUCAUAAUAGCGGCGAGGAAAAACA ....((.((((((((((((...)))))...((((.((...(((.(((..(((((((....))-----)))))..)).).)))...)).)))).)))))))))....... ( -29.60) >DroSec_CAF1 87350 97 - 1 -----------CGUUCCAUUUUAUGGAACAGUUAGGAAAACAGAGAUGAUAAGUGCAAUCGCACUGCACUUGAAAUACACUGCACUCAUAAUGGCGGCGAGGAA-AACA -----------.(((((((...))))))).(((................((((((((.......)))))))).....(.((((.(.......))))).).....-))). ( -22.20) >DroSim_CAF1 88782 109 - 1 AAUCCCGUAGUCGUUCCAUUUUAUGGAACCGUUAGGAAAACAGAGAUGAUAAGUGCAUUCGCACUGCACUUGAAAUACAUUGCACUCAUAAUAGCGGCGAGGAAAAACA ..((((((....(((((((...)))))))((((((.....).(((......(((((....)))))(((..((.....)).))).)))....)))))))).)))...... ( -28.30) >DroEre_CAF1 90384 98 - 1 CCGCC---CGCCGUUCCAUUUUAUGGAACCGUUAGAAAAACAGAGAUGAUAAGUGCAUUCGCACUGCAUUUCAAGU-------ACUCAUAAUAGCGGCGAG-AAAAGCA .((((---.((.(((((((...))))))).............((((((...(((((....))))).))))))....-------..........))))))..-....... ( -29.70) >DroYak_CAF1 92519 98 - 1 C-GGC---CGCCGCUCCAUAUUAUGGAACCGUUAGGAAAACAGAGAUGAUAAGUGCAUUCGCACUGCACUUUAGAU-------ACUCAUAAUAGCGGCGAGGAAAAACA .-...---(((((((....(((((((..((....)).....((((.((...(((((....))))).))))))....-------..)))))))))))))).......... ( -28.60) >consensus A_GCC___CGCCGUUCCAUUUUAUGGAACCGUUAGGAAAACAGAGAUGAUAAGUGCAUUCGCACUGCACUUGAAAUACA_UGCACUCAUAAUAGCGGCGAGGAAAAACA ........(((((((.....((((((....(((.....))).(((.((...(((((....))))).)))))..............)))))).))))))).......... (-15.44 = -16.32 + 0.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:34:08 2006