
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,459,947 – 4,460,100 |
| Length | 153 |
| Max. P | 0.715532 |

| Location | 4,459,947 – 4,460,066 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 91.47 |
| Mean single sequence MFE | -24.22 |
| Consensus MFE | -19.26 |
| Energy contribution | -19.26 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715532 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4459947 119 - 23771897 UAUCAUCCAGAUGUGACAUUUAAUCUGCAAGUGAUUUUUUGUGUCACUUCUGGUUAAAAACCUCUCACCAAUUGUUUCCAAAAUUAAACAUGGACAUGCAAGGGAUGUCCGAAUAAAUC ......(((((.(((((((..((((.......))))....))))))).)))))...................(((((........)))))(((((((.(...).)))))))........ ( -28.60) >DroSec_CAF1 37179 118 - 1 UAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGACU-UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCCAAAAUUAAACAUGGUCAUUAAAGCGAUGUCCGAAUAAAUC ...((((((((.(((((((..((((.......)).)-)..))))))).))))..............((((..(((((........))))))))).........))))............ ( -21.80) >DroSim_CAF1 37168 118 - 1 UAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGACU-UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCCAAAAUUAAACAUGGACAUUCAAGCGAUGUCCGAAUAAAUC .......((((.(((((((..((((.......)).)-)..))))))).))))....................(((((........)))))((((((((.....))))))))........ ( -24.50) >DroEre_CAF1 40084 117 - 1 UAUCAUUCAGAUGUGAUAUUUGAUCUGCAACUGAUU-UUUGUGUCACUUCUGUU-AAAAAGCUCUCAUCAAUUGCUUCGAAAAUUAAACAUGGACAUGCAUGGGAUGUUUGAAUAAAUC .......((((.(((((((..((((.......))))-...))))))).))))..-....(((...........)))(((((.(((...((((......)))).))).)))))....... ( -23.40) >DroYak_CAF1 39041 118 - 1 UAUCAUCCAGAUGUGUCAUUUGAUCUGCAACUGAUU-UUUGUGUCACUUCUGUUCAAAAACCUCUCACCAAUUGUUUCGAAAAUUAAACAUGGACAUGCAUGGGAUGUUUGAAUAAAUC .......((((.(((.(((..((((.......))))-...))).))).))))((((((....((((((((..(((((........)))))))).......)))))..))))))...... ( -22.81) >consensus UAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGAUU_UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCCAAAAUUAAACAUGGACAUGCAAGGGAUGUCCGAAUAAAUC ..((...((((.(((((((..((((.......))))....))))))).)))).......................................((((((.......))))))))....... (-19.26 = -19.26 + -0.00)


| Location | 4,459,987 – 4,460,100 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 81.35 |
| Mean single sequence MFE | -17.78 |
| Consensus MFE | -14.18 |
| Energy contribution | -14.46 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.628665 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4459987 113 - 23771897 -----CAGCACACUUAAUCCCAGCUUAAGCAUUUUUCAAUAUCAUCCAGAUGUGACAUUUAAUCUGCAAGUGAUUUUUUGUGUCACUUCUGGUUAAAAACCUCUCACCAAUUGUUUCC -----.((((............((....))...............(((((.(((((((..((((.......))))....))))))).)))))...................))))... ( -21.50) >DroSec_CAF1 37219 117 - 1 UCCCACAGCACACUUAAUCACAGCUUAAGCAUUUUUCAAUAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGACU-UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCC ...................((((.....(((....((((..((((......))))...))))..)))(((((((.-.....))))))))))).......................... ( -17.20) >DroSim_CAF1 37208 86 - 1 -------------------------------UUUUUCAAUAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGACU-UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCC -------------------------------...............((((.(((((((..((((.......)).)-)..))))))).))))........................... ( -14.80) >DroEre_CAF1 40124 116 - 1 GCCCACAGCAGACUUAAUCCCAACUUAAGCAUUUUUCUAUAUCAUUCAGAUGUGAUAUUUGAUCUGCAACUGAUU-UUUGUGUCACUUCUGUU-AAAAAGCUCUCAUCAAUUGCUUCG ......(((((.(((((.......))))).................((((.(((((((..((((.......))))-...))))))).))))..-................)))))... ( -19.90) >DroYak_CAF1 39081 117 - 1 GCCCACAGCACACUUAAUCCCAAUUUAAGCAUUUUUCUAUAUCAUCCAGAUGUGUCAUUUGAUCUGCAACUGAUU-UUUGUGUCACUUCUGUUCAAAAACCUCUCACCAAUUGUUUCG ......((((..(((((.......))))).................((((.(((.(((..((((.......))))-...))).))).))))....................))))... ( -15.50) >consensus _CCCACAGCACACUUAAUCCCAACUUAAGCAUUUUUCAAUAUCAUCCAGAUGUGACAUUUGAUCUGCAAGUGAUU_UUUGUGUCACUUCUGUUAAAAAACCUCUCACCAAUUGUUUCC ..............................................((((.(((((((..((((.......))))....))))))).))))........................... (-14.18 = -14.46 + 0.28)
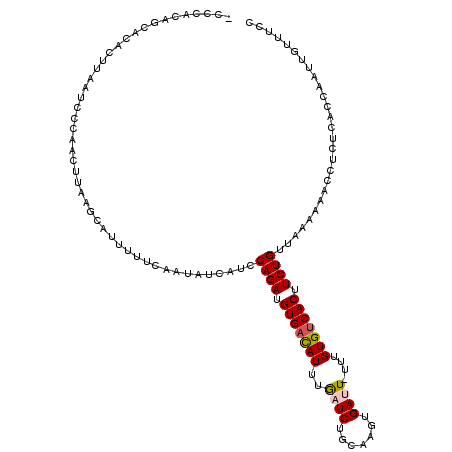



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:14 2006