
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,341,513 – 4,341,623 |
| Length | 110 |
| Max. P | 0.954903 |

| Location | 4,341,513 – 4,341,623 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 81.57 |
| Mean single sequence MFE | -30.77 |
| Consensus MFE | -20.20 |
| Energy contribution | -20.23 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.954903 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
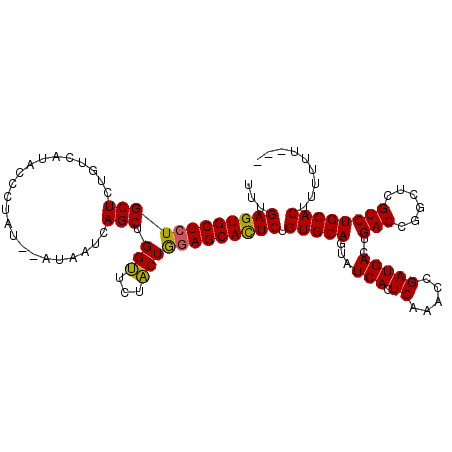
>3L_DroMel_CAF1 4341513 110 + 23771897 AUGAAAAAAGUGCAGACGAGCCGGUCAGUCAUCGGUUUGAGUGAUACUGCACAGAAUGCUCCAGUAGAACCAGCUGAUUAG--AUAGGCUGUGACAGAGCAGAGCACUCAAA .........((((((.(((((((((.....)))))))))(....).)))))).((.(((((((((.......)))).....--....(((.......))).))))).))... ( -32.60) >DroSim_CAF1 27754 96 + 1 ---AAAAAAGUGCA-------------GUCAUCGGUUUGAGUGAUACUGCACAGAGUGCUCCAGUAGAACCAGCUGAUUAUAUAUAGGGUAUGACAGAGCAGAGCACUCAAA ---......(((((-------------((.((((.......))))))))))).((((((((((((.......)))).((((((.....)))))).......))))))))... ( -31.90) >DroYak_CAF1 27445 106 + 1 ----AAAAAGUGCAGACGAGCCGGUCAGUCAUCGGUAUGAAUGAUACUGCACAGAGUGCUCUAGAAAAUCCAGCUGAUUAC--AUAGGGUAUGACAGAGCACAGCACUCAAA ----....(((((......((..((((.((((....)))).))))...)).....(((((((.....((((..........--...)))).....))))))).))))).... ( -27.82) >consensus ___AAAAAAGUGCAGACGAGCCGGUCAGUCAUCGGUUUGAGUGAUACUGCACAGAGUGCUCCAGUAGAACCAGCUGAUUAC__AUAGGGUAUGACAGAGCAGAGCACUCAAA .........((((((((......))).(((((........)))))..))))).((((((((((((.......))))..(((.......)))..........))))))))... (-20.20 = -20.23 + 0.04)



| Location | 4,341,513 – 4,341,623 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 81.57 |
| Mean single sequence MFE | -27.30 |
| Consensus MFE | -17.92 |
| Energy contribution | -17.62 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.771131 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4341513 110 - 23771897 UUUGAGUGCUCUGCUCUGUCACAGCCUAU--CUAAUCAGCUGGUUCUACUGGAGCAUUCUGUGCAGUAUCACUCAAACCGAUGACUGACCGGCUCGUCUGCACUUUUUUCAU ((((((((..((((.(.....((((....--.......))))((((.....)))).....).))))...))))))))..(((((((....)).))))).............. ( -28.00) >DroSim_CAF1 27754 96 - 1 UUUGAGUGCUCUGCUCUGUCAUACCCUAUAUAUAAUCAGCUGGUUCUACUGGAGCACUCUGUGCAGUAUCACUCAAACCGAUGAC-------------UGCACUUUUUU--- ...(((((((((.....((...(((................)))...)).))))))))).((((((((((.........))).))-------------)))))......--- ( -26.19) >DroYak_CAF1 27445 106 - 1 UUUGAGUGCUGUGCUCUGUCAUACCCUAU--GUAAUCAGCUGGAUUUUCUAGAGCACUCUGUGCAGUAUCAUUCAUACCGAUGACUGACCGGCUCGUCUGCACUUUUU---- ...(((((((.....(((.((((...)))--)....)))((((.....))))))))))).(((((((((.....)))).(((((((....)).)))))))))).....---- ( -27.70) >consensus UUUGAGUGCUCUGCUCUGUCAUACCCUAU__AUAAUCAGCUGGUUCUACUGGAGCACUCUGUGCAGUAUCACUCAAACCGAUGACUGACCGGCUCGUCUGCACUUUUUU___ ...((((((((((((......................))).(((...)))))))))))).(((((...(((.((.....)))))..(((......))))))))......... (-17.92 = -17.62 + -0.30)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:27 2006