
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,337,943 – 4,338,058 |
| Length | 115 |
| Max. P | 0.555364 |

| Location | 4,337,943 – 4,338,058 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -30.55 |
| Consensus MFE | -23.84 |
| Energy contribution | -26.15 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4337943 115 + 23771897 ACUGCAAUUUCAAGUGAUUUGUGGCCGAUUUGUGUUGCCGAUUUCUUGCAAAAUGGCACACACUCCCUCUGGAAACCUAAAUCCACUCGAAACGAGAGCUCUCCAGGCAUUAACU ..(((..((((.(((((((((.((......(((((.((((.(((.....))).))))))))).(((....)))..))))))).)))).)))).(((....)))...)))...... ( -26.20) >DroSec_CAF1 24161 102 + 1 ACUGCAAUUUCAAGUGAUUUGUGGCCG-------------AUUUCUGGCAAAAUGGCACACACUCCCGCUGGAAACCCAGAUCCACUCGAAAGGAGAGCUCUCCAGGCAUUAACU ..(((..((((.((((...((((((((-------------.(((.....))).)))).))))......((((....))))...)))).))))((((....))))..)))...... ( -29.40) >DroSim_CAF1 24209 115 + 1 ACUGCAAUUUCAAGUGAUUUGUGGCCGAUUUGUGUUGCCGAUUUCUGGCAAAAUGGCACACACUCCCGCUGGAAACCCAGAUCCACUCGAAAGGAGAGCUCUCCAGGCAUUAACU ..(((..((((.((((...((((((((.......((((((.....))))))..)))).))))......((((....))))...)))).))))((((....))))..)))...... ( -35.60) >DroYak_CAF1 23708 115 + 1 ACUGCAAUUUCAAGUGAUUUGUGGCCGAUUUUUGUUGCCGAUUUCUGGCAAAAUGGCACACUCUCCCUCUGCAAAUCCAGAUCCACUCGAAAGGAGGGCUCUCCAGGCAUUAACU ..(((..((((.((((....(((((((.......((((((.....))))))..)))).)))......((((......))))..)))).))))((((....))))..)))...... ( -31.00) >consensus ACUGCAAUUUCAAGUGAUUUGUGGCCGAUUUGUGUUGCCGAUUUCUGGCAAAAUGGCACACACUCCCGCUGGAAACCCAGAUCCACUCGAAAGGAGAGCUCUCCAGGCAUUAACU ..(((..((((.((((....(((((((.......((((((.....))))))..)))).))).......((((....))))...)))).))))((((....))))..)))...... (-23.84 = -26.15 + 2.31)

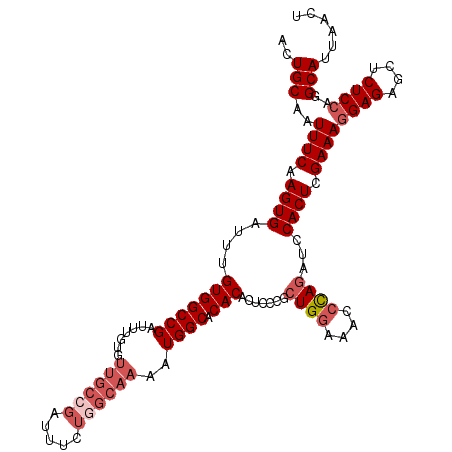

| Location | 4,337,943 – 4,338,058 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -34.88 |
| Consensus MFE | -26.26 |
| Energy contribution | -26.82 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4337943 115 - 23771897 AGUUAAUGCCUGGAGAGCUCUCGUUUCGAGUGGAUUUAGGUUUCCAGAGGGAGUGUGUGCCAUUUUGCAAGAAAUCGGCAACACAAAUCGGCCACAAAUCACUUGAAAUUGCAGU ......(((...(((....)))((((((((((.((((.(((((((....)))((((.(((((((((....))))).)))))))).....))))..)))))))))))))).))).. ( -33.70) >DroSec_CAF1 24161 102 - 1 AGUUAAUGCCUGGAGAGCUCUCCUUUCGAGUGGAUCUGGGUUUCCAGCGGGAGUGUGUGCCAUUUUGCCAGAAAU-------------CGGCCACAAAUCACUUGAAAUUGCAGU ......(((..((((....))))((((((((((..((((....))))......((((.((((((((....)))))-------------.)))))))..))))))))))..))).. ( -33.20) >DroSim_CAF1 24209 115 - 1 AGUUAAUGCCUGGAGAGCUCUCCUUUCGAGUGGAUCUGGGUUUCCAGCGGGAGUGUGUGCCAUUUUGCCAGAAAUCGGCAACACAAAUCGGCCACAAAUCACUUGAAAUUGCAGU ......(((..((((....))))((((((((((..((((....))))(((...(((((((((((((....))))).))).)))))..)))........))))))))))..))).. ( -37.60) >DroYak_CAF1 23708 115 - 1 AGUUAAUGCCUGGAGAGCCCUCCUUUCGAGUGGAUCUGGAUUUGCAGAGGGAGAGUGUGCCAUUUUGCCAGAAAUCGGCAACAAAAAUCGGCCACAAAUCACUUGAAAUUGCAGU ......(((..((((....))))((((((((((.((((......))))......(((.((((((((((((....).))))....)))).))))))...))))))))))..))).. ( -35.00) >consensus AGUUAAUGCCUGGAGAGCUCUCCUUUCGAGUGGAUCUGGGUUUCCAGAGGGAGUGUGUGCCAUUUUGCCAGAAAUCGGCAACACAAAUCGGCCACAAAUCACUUGAAAUUGCAGU ......(((..((((....))))((((((((((........((((....)))).(((.((((((((....))))).(....).......))))))...))))))))))..))).. (-26.26 = -26.82 + 0.56)

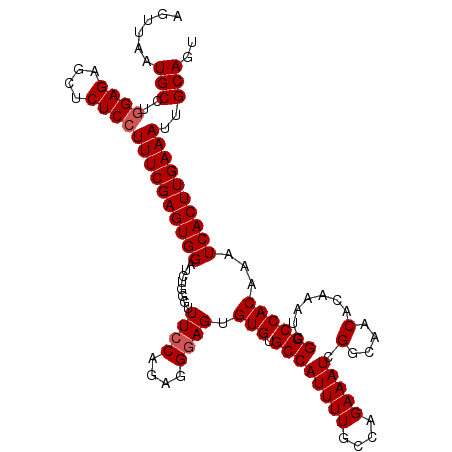
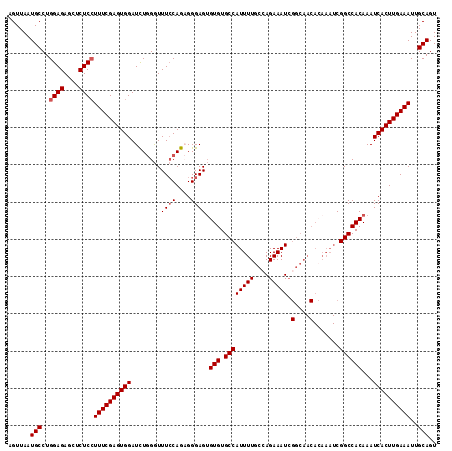
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:25 2006