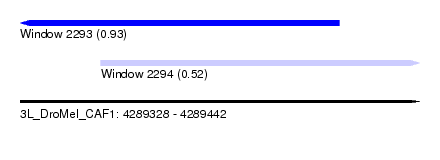

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,289,328 – 4,289,442 |
| Length | 114 |
| Max. P | 0.927848 |

| Location | 4,289,328 – 4,289,419 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.45 |
| Mean single sequence MFE | -22.67 |
| Consensus MFE | -14.23 |
| Energy contribution | -14.23 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.927848 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4289328 91 - 23771897 CGCCGCCUUGUGCAAUUUUCACGCAGUUUUU-CAUGGCAGCGUGAAAAUGU--------CAAACAA-CAACGGGAUAG---------AAAGAAA------AAAGAAAAGC----GACCGA ..(((..((((...((((((((((.(((...-...))).))))))))))..--------...))))-...))).....---------.......------..........----...... ( -20.50) >DroPse_CAF1 13661 99 - 1 CGCCGCCUUCUGCAAUUUUCACGCAGUUUUUCCAUGGCAGCGUGAAAAUGU--------CAGGCAGGCAA----ACAG---------AAAGAAAAAAUUGAAAGAAAAACCAGAAAACAA .(((((((...(((.(((((((((.(((.......))).))))))))))))--------.)))).)))..----....---------................................. ( -25.90) >DroSec_CAF1 12881 91 - 1 CGCCGCCUUGUGCAAUUUUCACGCAGUUUUU-CAUGGCAGCGUGAAAAUGU--------CAAACAA-CAACAGGAUAG---------AAAGAAA------AAAGAAAAGC----GACCAA ..((...((((...((((((((((.(((...-...))).))))))))))..--------...))))-.....))....---------.......------..........----...... ( -18.00) >DroWil_CAF1 13649 96 - 1 CGCCACCU-AAGCAAUUUUCACGCAGUUUUU-CAUGGCAGCGUGAAAAUGUUAACAAAGCAAACAA-CAGCAGCAAGG---------CAGGCAA------CAACAACAAC----GAC--A .(((.(((-..((.((((((((((.(((...-...))).)))))))))).........((......-..)).)).)))---------..)))..------..........----...--. ( -24.80) >DroYak_CAF1 13193 91 - 1 CGCCGCCUUGUGCAAUUUUCACGCAGUUUUU-CAUGGCAGCGUGAAAAUGU--------CAAACAA-CAACGGGCUAG---------AAAGAAA------AUAGAAAAGC----GACCGA ..(((..((((...((((((((((.(((...-...))).))))))))))..--------...))))-...)))(((..---------.......------.......)))----...... ( -21.19) >DroAna_CAF1 12803 103 - 1 CGCCGCCUUGUGCAAUUUUCACGCAGUUUUU-CAUGGCAGCGUGAAAAUGU--------CAAACAA-CAACGGCAUGGCUUAAAAAAAAAGAAA---CAGAAAGAAAAGC----GUCCAA .((((..((((...((((((((((.(((...-...))).))))))))))..--------...))))-...))))...((((.............---.........))))----...... ( -25.65) >consensus CGCCGCCUUGUGCAAUUUUCACGCAGUUUUU_CAUGGCAGCGUGAAAAUGU________CAAACAA_CAACGGGAUAG_________AAAGAAA______AAAGAAAAGC____GACCAA .((........)).((((((((((.(((.......))).))))))))))....................................................................... (-14.23 = -14.23 + -0.00)

| Location | 4,289,351 – 4,289,442 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 85.86 |
| Mean single sequence MFE | -26.55 |
| Consensus MFE | -19.59 |
| Energy contribution | -19.48 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.518006 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4289351 91 + 23771897 --CUAUCCCGUUG-UUGUUUGACAUUUUCACGCUGCCAUG-AAAAACUGCGUGAAAAUUGCACAAGGCGGCG-AGACAAUUGCCGUUAAUUU--GAAA --........(((-(.((.....((((((((((.......-.......)))))))))).))))))(((((((-(.....)))))))).....--.... ( -24.04) >DroPse_CAF1 13694 89 + 1 --CUGU----UUGCCUGCCUGACAUUUUCACGCUGCCAUGGAAAAACUGCGUGAAAAUUGCAGAAGGCGGCG-AGACAAUUGCCGUUAAUUU--GAGA --....----..((((..(((..((((((((((...............))))))))))..))).))))((((-(.....)))))........--.... ( -29.86) >DroGri_CAF1 14287 83 + 1 --CUGU------G-UUUU-----GUGUUCACGCUGUGUUGGAAAAACUGCGUGAAAAUUGCACA-GGCGGCAAAGACAAUUGCCGUUAAUUUUAAAAA --((((------(-(..(-----...(((((((...(((.....))).))))))).)..)))))-)(((((((......)))))))............ ( -30.10) >DroSec_CAF1 12904 91 + 1 --CUAUCCUGUUG-UUGUUUGACAUUUUCACGCUGCCAUG-AAAAACUGCGUGAAAAUUGCACAAGGCGGCG-AGACAAUUGCCGUUAAUUU--GAAA --........(((-(.((.....((((((((((.......-.......)))))))))).))))))(((((((-(.....)))))))).....--.... ( -24.04) >DroYak_CAF1 13216 91 + 1 --CUAGCCCGUUG-UUGUUUGACAUUUUCACGCUGCCAUG-AAAAACUGCGUGAAAAUUGCACAAGGCGGCG-AGACAAUUGCCGUUAAUUU--GAAA --...(((..(((-(.((.....((((((((((.......-.......)))))))))).)))))))))((((-(.....)))))........--.... ( -24.94) >DroAna_CAF1 12836 93 + 1 AGCCAUGCCGUUG-UUGUUUGACAUUUUCACGCUGCCAUG-AAAAACUGCGUGAAAAUUGCACAAGGCGGCG-AGACAAUUGCCGUUAAUUU--GAAA ......(((.(((-(.((.....((((((((((.......-.......)))))))))).)))))))))((((-(.....)))))........--.... ( -26.34) >consensus __CUAUCCCGUUG_UUGUUUGACAUUUUCACGCUGCCAUG_AAAAACUGCGUGAAAAUUGCACAAGGCGGCG_AGACAAUUGCCGUUAAUUU__GAAA .......................((((((((((...............)))))))))).......(((((((........)))))))........... (-19.59 = -19.48 + -0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:35 2006