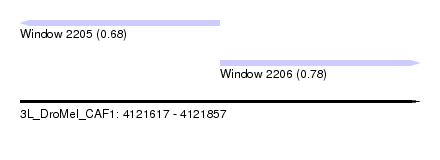
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 4,121,617 – 4,121,857 |
| Length | 240 |
| Max. P | 0.783055 |

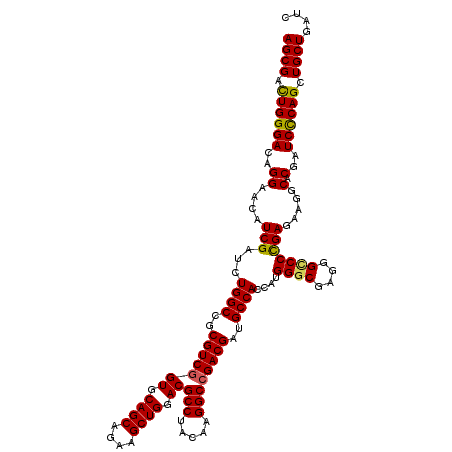
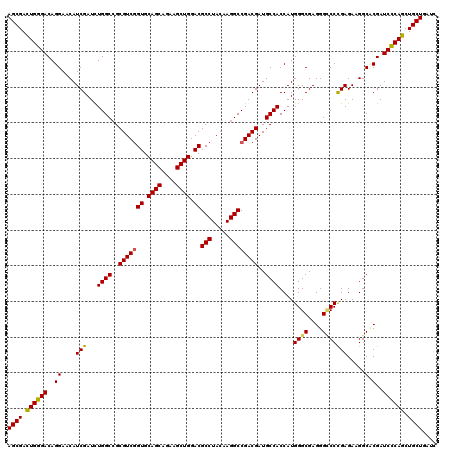
| Location | 4,121,617 – 4,121,737 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.56 |
| Mean single sequence MFE | -51.15 |
| Consensus MFE | -46.69 |
| Energy contribution | -46.50 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.679288 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4121617 120 - 23771897 AGCGACUGGGACAGGAACAUCGAUUUGGCCGCGUCGGUGCAGCAGAAGCUGGACGCCUACAAGGCCGACGAUGCCACCAUGGGCGAGGGCCCCGAGAAGGCACGAUCCCAGCUGCUGAUC ((((.((((((..((....(((...((((..(((((((.((((....)))).))(((.....))))))))..))))....((((....))))))).....).)..)))))).)))).... ( -51.60) >DroSec_CAF1 3236 120 - 1 AGCGAUUGGGACAGGAACAUCGAUCUGGCCGCGUCUGUGCAGCAGAAGCUGGACGCCUACAAGGCCGACGAUGCCACCAUGGGCGAGGGCCCUGAGAAGGCACGAUCCCAGCUGCUGAUU ((((.((((((..(....((((.((.(((((((((....((((....)))))))))......))))))))))(((.(((.((((....)))))).)..))).)..)))))).)))).... ( -47.20) >DroEre_CAF1 3432 120 - 1 AGCGACUGGGACAGGAACAUCGAUUUGGCCGCGUCGGUGCAGCAGAAGCUGGACGCCUACAAGGCCGACGAUGCCACCAUGGGCGAGGGUCCCGAGAAGGCCCGAUCCCAGCUGCUGAUC ((((.((((((..((..(.(((...((((..(((((((.((((....)))).))(((.....))))))))..))))....((((....)))))))...)..))..)))))).)))).... ( -52.60) >DroYak_CAF1 3882 120 - 1 AGCGACUGGGACAGGAACAUCGAUCUGGCGGCGUCGGUGCAGCAGAAGCUGGACGCCUACAAGGCCGACGAUGCCACCAUGGGCGAGGGACCCGAGAAGGCUCGAUCUCAGCUGCUGAUC ((((.((((((..((..(.(((..((((.(((((((((.((((....)))).))(((.....)))...))))))).))).)..))).)..))((((....)))).)))))).)))).... ( -53.20) >consensus AGCGACUGGGACAGGAACAUCGAUCUGGCCGCGUCGGUGCAGCAGAAGCUGGACGCCUACAAGGCCGACGAUGCCACCAUGGGCGAGGGCCCCGAGAAGGCACGAUCCCAGCUGCUGAUC ((((.((((((..((....(((...((((..(((((((.((((....)))).))(((.....))))))))..))))....((((....))))))).....).)..)))))).)))).... (-46.69 = -46.50 + -0.19)

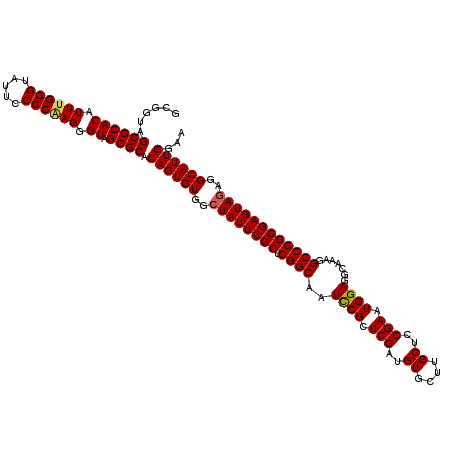
| Location | 4,121,737 – 4,121,857 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.50 |
| Mean single sequence MFE | -48.75 |
| Consensus MFE | -47.34 |
| Energy contribution | -47.52 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.783055 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 4121737 120 + 23771897 GCGGUAGCGCACAUUUGGGUACUCCCCAAGGGUAGCGCACAGUGUGGCUAUUUGCUCGGCAAUUCGCUCGAUGUGCUUGCUCCGAAUGGAGGCAAAGGCCGGCGAGUAGAGGCAUUGGAA ......((((((.((((((.....)))))).)).)))).((((((..((((((((.((((.....((.......))(((((((....))).))))..))))))))))))..))))))... ( -51.00) >DroSec_CAF1 3356 120 + 1 GCGGUAGCGCACAUUUGGGUAUUCCCCAAGGGUAGCGCACAGUGUGGCUAUUUGCUCGGCAAUCCGCUCGAUGUGCUUGCUCCGAAUGGAGGCAAAGGCCGGCGAGUAGAGGCAUUGGAA ......((((((.((((((.....)))))).)).)))).((((((..((((((((.((((....(((.....))).(((((((....))).))))..))))))))))))..))))))... ( -51.00) >DroEre_CAF1 3552 120 + 1 GCGGUAGCGCACAUUUGGGUAUUCCCCGAGGGUAGCGCACAGUGUGGCUAUUUGCUCGGCAAUCCGCUCGAUGUGCUUGCUCCGAAUGGAGGCAAAGGCCGGCGAGUAUAGGCAUUGGAA ......((((((.((((((.....)))))).)).)))).((((((...(((((((.((((....(((.....))).(((((((....))).))))..)))))))))))...))))))... ( -45.30) >DroYak_CAF1 4002 120 + 1 GCGGUAGCGCACAUUCGGGUAUUCCCCAAGGGUAGCGCACAGUGUGGCUAUUUGCUCGGCAAUCCGCUCGAUGUGCUUGCUGCGAAUGGAGGCAAAGGCCGGCGAGUAGAGGCAUUGGAA ......((((((.((.(((....))).))..)).)))).((((((..((((((((.((((......(((.((.(((.....))).)).)))......))))))))))))..))))))... ( -47.70) >consensus GCGGUAGCGCACAUUUGGGUAUUCCCCAAGGGUAGCGCACAGUGUGGCUAUUUGCUCGGCAAUCCGCUCGAUGUGCUUGCUCCGAAUGGAGGCAAAGGCCGGCGAGUAGAGGCAUUGGAA ......((((((.((((((.....)))))).)).)))).((((((..((((((((.((((..((((.(((..((....))..))).)))).......))))))))))))..))))))... (-47.34 = -47.52 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:07:02 2006