
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,995,977 – 3,996,068 |
| Length | 91 |
| Max. P | 0.963688 |

| Location | 3,995,977 – 3,996,068 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 77.87 |
| Mean single sequence MFE | -26.25 |
| Consensus MFE | -16.33 |
| Energy contribution | -16.82 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919163 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
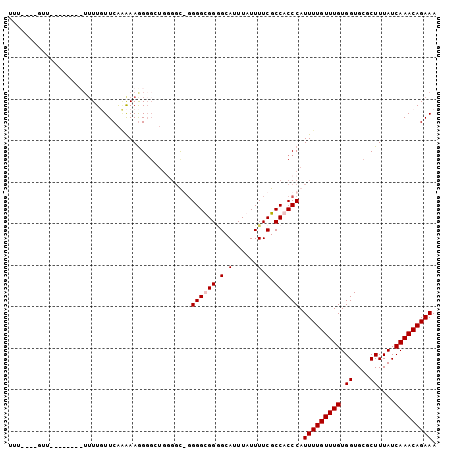
>3L_DroMel_CAF1 3995977 91 + 23771897 UUU----CUU--------UUUUUUUCAAAAAGGGGCUGUUGCGUGGGCGGGGCAUUUAUUUUCGCCACCCAUUUUGUUUGUGGUGCGCUUUAUCAAACAGAAA ...----...--------.............(((((....))...((((..(.......)..)))).))).(((((((((.((....))....))))))))). ( -23.50) >DroSec_CAF1 68577 89 + 1 UUU----GUU----------UUUUUCAAAAAGGGGCUGUGCCGGGGGCGGGGCAUUUAUUUUCGCCACCCAUUUUGUUUGUGGUGCGCUUUAUCAAACAGAAA ...----...----------.............(((...)))(((((((..(.......)..)))).))).(((((((((.((....))....))))))))). ( -25.40) >DroEre_CAF1 71746 98 + 1 UUUUGUUGUUUUUUUCAUUUUUGUUCGGGAAAG----UGGGU-GGGGUGGCGCAUUUAUUUUCGCCACCCAUUUUGUUUGUGGUGCGCUUUAUCAAACAGAAA .............(((((((((......)))))----)))).-.((((((((.(......).)))))))).(((((((((.((....))....))))))))). ( -28.40) >DroYak_CAF1 73337 97 + 1 -----UUUUUUUUUAUAUUUUUGUUCACAAAGGGGAUGGGGU-GGGGUGGAGCAUUUAUUUUCGCCACCCAUUUUGUUUGUGGUGCGCUUUAUCAAACAGAAA -----............((((((((((((((.((((((((..-..(((((((.......))))))).)))))))).))))))(....).......)))))))) ( -27.70) >consensus UUU____GUU________UUUUGUUCAAAAAGGGGCUGGGGC_GGGGCGGGGCAUUUAUUUUCGCCACCCAUUUUGUUUGUGGUGCGCUUUAUCAAACAGAAA ............................................((((((.(.(......).).)))))).(((((((((.((....))....))))))))). (-16.33 = -16.82 + 0.50)


| Location | 3,995,977 – 3,996,068 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 77.87 |
| Mean single sequence MFE | -17.39 |
| Consensus MFE | -12.97 |
| Energy contribution | -12.72 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.963688 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
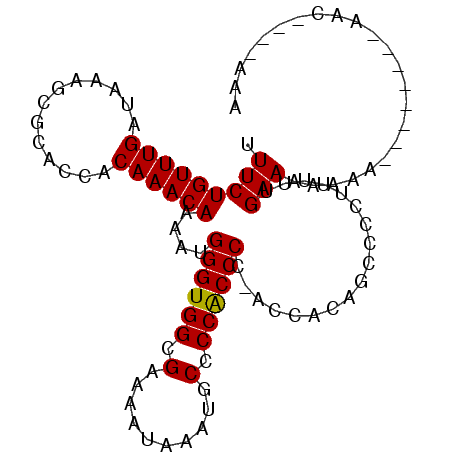
>3L_DroMel_CAF1 3995977 91 - 23771897 UUUCUGUUUGAUAAAGCGCACCACAAACAAAAUGGGUGGCGAAAAUAAAUGCCCCGCCCACGCAACAGCCCCUUUUUGAAAAAAA--------AAG----AAA ....((((((.............))))))...(((((((.(...........)))))))).((....))..((((((.....)))--------)))----... ( -16.82) >DroSec_CAF1 68577 89 - 1 UUUCUGUUUGAUAAAGCGCACCACAAACAAAAUGGGUGGCGAAAAUAAAUGCCCCGCCCCCGGCACAGCCCCUUUUUGAAAAA----------AAC----AAA ((((((((((.............))))))....(((.((((.............)))))))(((...))).......))))..----------...----... ( -17.34) >DroEre_CAF1 71746 98 - 1 UUUCUGUUUGAUAAAGCGCACCACAAACAAAAUGGGUGGCGAAAAUAAAUGCGCCACCCC-ACCCA----CUUUCCCGAACAAAAAUGAAAAAAACAACAAAA ....((((((..((((.(.....).........((((((((..........)))))))).-.....----))))..))))))..................... ( -20.10) >DroYak_CAF1 73337 97 - 1 UUUCUGUUUGAUAAAGCGCACCACAAACAAAAUGGGUGGCGAAAAUAAAUGCUCCACCCC-ACCCCAUCCCCUUUGUGAACAAAAAUAUAAAAAAAAA----- .....((((....))))....((((((......((((((.(................)))-)))).......))))))....................----- ( -15.31) >consensus UUUCUGUUUGAUAAAGCGCACCACAAACAAAAUGGGUGGCGAAAAUAAAUGCCCCACCCC_ACCACAGCCCCUUUUUGAAAAAAA________AAC____AAA .(((((((((.............))))))....((((((.(..........).))))))..................)))....................... (-12.97 = -12.72 + -0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:05:54 2006