
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,990,181 – 3,990,331 |
| Length | 150 |
| Max. P | 0.841685 |

| Location | 3,990,181 – 3,990,300 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.97 |
| Mean single sequence MFE | -29.22 |
| Consensus MFE | -19.74 |
| Energy contribution | -20.73 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.841685 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3990181 119 + 23771897 -ACUGUUCCAAGGACUUUAAAAACCAUACUAAAAGGACAUGAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGGUCAAAUUAUUGAGUUCCUGGUUGAGCACAAAA -..(((((...(..((((.............))))..).((.(((((........))))).)).......(((((((((.((((.........)))).)))))))))...)))))..... ( -22.52) >DroSec_CAF1 62544 120 + 1 GACUGUUCCAAGGACUUUAAAGGCCAUGCUUAAAGGACAUGAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGGCCAACUUAUUGAGUUCCUGGGUGAGCAGAAAA ..(((((((..(..(((((..(((...))))))))..).((.(((((........))))).)).......(((((((((.....(((.....)))...))))))))).).)))))).... ( -31.70) >DroEre_CAF1 64071 120 + 1 GACUUUUCCAAGGACUUUGCAAGCCUUGCUUAAAGGCCAAGAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUGCGUUGAAUACAAACUUAUAGAGUUCCUGUGCGAGCACAAAA .................((((((((((.....)))))((((............)))).)))))((....(((((((((.(..(((........)))..))))))))))....))...... ( -27.60) >DroYak_CAF1 66702 120 + 1 GACUACUCCAAGGACUUUAAAAGCCUUGCUUAAAGGACAAGGGAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCACACAAUACAAACUUAUGGAGUUCCUGUGCGAGCACAAGA ......(((..(..(((((..(((...))))))))..)...))).........(((((..((..((...((((((((((((...............)))))))))))).)).))))))). ( -35.06) >consensus GACUGUUCCAAGGACUUUAAAAGCCAUGCUUAAAGGACAAGAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGACAAACUUAUUGAGUUCCUGGGCGAGCACAAAA ...........(..(((((..(((...))))))))..)..(.(((((........))))).).((....((((((((((...................))))))))))....))...... (-19.74 = -20.73 + 1.00)

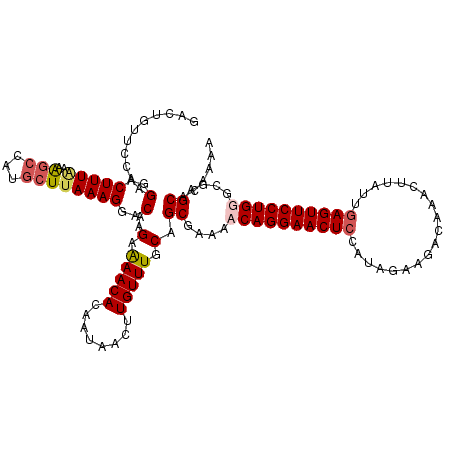
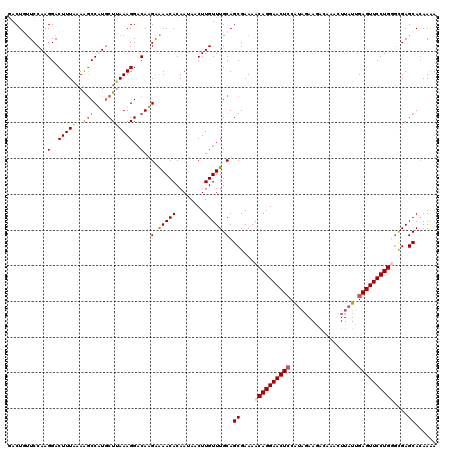
| Location | 3,990,220 – 3,990,331 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 88.55 |
| Mean single sequence MFE | -27.17 |
| Consensus MFE | -21.56 |
| Energy contribution | -22.56 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.753402 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3990220 111 + 23771897 GAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGGUCAAAUUAUUGAGUUCCUGGUUGAGCACAAAACCUCCUGUUACUGGGCAUAGCAAAGUCGUAA .........(..((((((((((.((..((.(((((((((.((((.........)))).))))))))).))..)).........((.......))))).))).))))..).. ( -25.60) >DroSec_CAF1 62584 111 + 1 GAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGGCCAACUUAUUGAGUUCCUGGGUGAGCAGAAAACCUCCUGUUACUGGGCAUAGCAAAGUCGUAA .........(..((((((((((........(((((((((.....(((.....)))...)))))))))(((((.(((........))))))))..))).))).))))..).. ( -28.80) >DroEre_CAF1 64111 109 + 1 GAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUGCGUUGAAUACAAACUUAUAGAGUUCCUGUGCGAGCACAAAAA--CCUGUUACUGGGCAUAGGAAAGUCGUAA ....((.((.((((..((((...((....(((((((((.(..(((........)))..))))))))))....))....)))--)..)))).))(((........))))).. ( -23.40) >DroYak_CAF1 66742 109 + 1 GGGAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCACACAAUACAAACUUAUGGAGUUCCUGUGCGAGCACAAGAA--CCUGUUACUGGGCAUAGCAAAGUCGUAA .........(..((((((((((.((....((((((((((((...............))))))))))))....)).......--((.......))))).))).))))..).. ( -30.86) >consensus GAAAACACAAUAACUUGUUUGCAGCGAAAACAGGAACUCCAUAGAAGACAAACUUAUUGAGUUCCUGGGCGAGCACAAAAA__CCUGUUACUGGGCAUAGCAAAGUCGUAA .........(..((((((((((.((....((((((((((...................))))))))))....)).........((.......))))).))).))))..).. (-21.56 = -22.56 + 1.00)

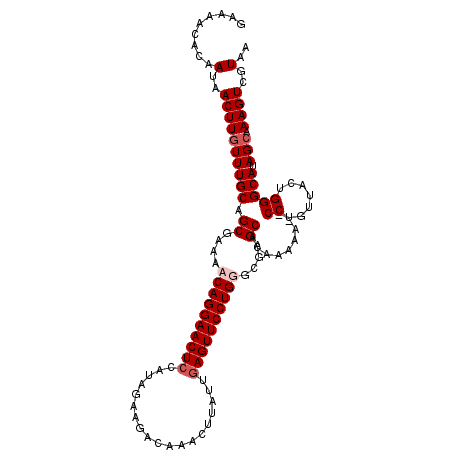

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:05:52 2006