
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,989,223 – 3,989,314 |
| Length | 91 |
| Max. P | 0.768304 |

| Location | 3,989,223 – 3,989,314 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 84.72 |
| Mean single sequence MFE | -31.32 |
| Consensus MFE | -26.09 |
| Energy contribution | -28.77 |
| Covariance contribution | 2.69 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.768304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3989223 91 + 23771897 AAUUUUGGGUUUAAGCACGUCUAGAGGUCGGAGCCUGCGGGCGACUAUGGCCAUGGCCAUGGG------UAUGGGUAUGGCUAUGAGUAUACUCUAG ......((((.((..(.(((((..((((....))))..)))))...(((((((((.((((...------.)))).))))))))))..)).))))... ( -32.50) >DroSec_CAF1 61604 97 + 1 AAUUUUGGGUUUAAGCACGUCUAGAGGUCGGAGCCUGCGGGCGACUAUGGCCAUGGCCAUGGGUUUGGGUGUGGGUAUGGCUAUGGGUAUACUCUAG ......((((....((.(((((..((((....))))..))))).(((((((((((.(((((........))))).)))))))))))))..))))... ( -35.60) >DroSim_CAF1 65171 97 + 1 AAUUUUGGGUUUAAGCACGUCUAGAGGUCGGAGCCUGCGGGCGACUAUGGCCAUGGCCAUGGGUUUGGGUGUGGGUAUGGCUAUGGGUAUACUCUAG ......((((....((.(((((..((((....))))..))))).(((((((((((.(((((........))))).)))))))))))))..))))... ( -35.60) >DroEre_CAF1 63119 79 + 1 AAUUUUGGGUUUAAGCACGUCUAGAGGUCGGAGCCUGCGGGCGAGCAUGGC------------------CAUGGGUAAGGCAAUGGGUAUACUCCGG .....((((.....((.(((((..((((....))))..))))).))(((.(------------------(((..(.....).)))).)))..)))). ( -21.60) >consensus AAUUUUGGGUUUAAGCACGUCUAGAGGUCGGAGCCUGCGGGCGACUAUGGCCAUGGCCAUGGG______UAUGGGUAUGGCUAUGGGUAUACUCUAG ......((((....((.(((((..((((....))))..))))).(((((((((((.(((((........))))).)))))))))))))..))))... (-26.09 = -28.77 + 2.69)


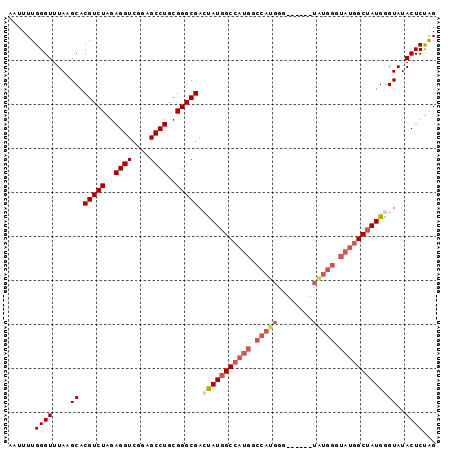
| Location | 3,989,223 – 3,989,314 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 84.72 |
| Mean single sequence MFE | -20.30 |
| Consensus MFE | -13.09 |
| Energy contribution | -14.40 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.700800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3989223 91 - 23771897 CUAGAGUAUACUCAUAGCCAUACCCAUA------CCCAUGGCCAUGGCCAUAGUCGCCCGCAGGCUCCGACCUCUAGACGUGCUUAAACCCAAAAUU ((((((..........(((((..((((.------...))))..)))))....((((((....)))...))))))))).................... ( -22.10) >DroSec_CAF1 61604 97 - 1 CUAGAGUAUACCCAUAGCCAUACCCACACCCAAACCCAUGGCCAUGGCCAUAGUCGCCCGCAGGCUCCGACCUCUAGACGUGCUUAAACCCAAAAUU (((((((((..((((.(((((................))))).))))..)))((((((....)))...))))))))).................... ( -21.49) >DroSim_CAF1 65171 97 - 1 CUAGAGUAUACCCAUAGCCAUACCCACACCCAAACCCAUGGCCAUGGCCAUAGUCGCCCGCAGGCUCCGACCUCUAGACGUGCUUAAACCCAAAAUU (((((((((..((((.(((((................))))).))))..)))((((((....)))...))))))))).................... ( -21.49) >DroEre_CAF1 63119 79 - 1 CCGGAGUAUACCCAUUGCCUUACCCAUG------------------GCCAUGCUCGCCCGCAGGCUCCGACCUCUAGACGUGCUUAAACCCAAAAUU .((((((.........(((........)------------------))..(((......))).))))))............................ ( -16.10) >consensus CUAGAGUAUACCCAUAGCCAUACCCACA______CCCAUGGCCAUGGCCAUAGUCGCCCGCAGGCUCCGACCUCUAGACGUGCUUAAACCCAAAAUU ((((((...............................(((((....)))))....(((....)))......)))))).................... (-13.09 = -14.40 + 1.31)
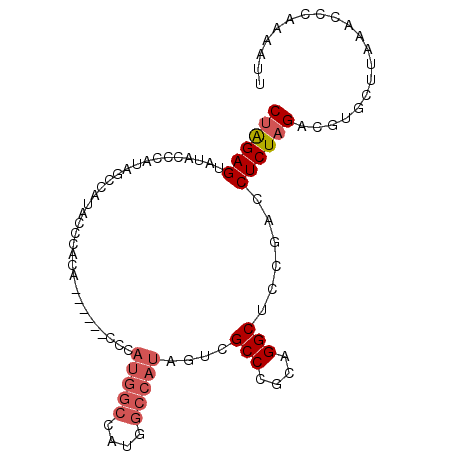



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:05:50 2006