
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,966,792 – 3,966,890 |
| Length | 98 |
| Max. P | 0.999818 |

| Location | 3,966,792 – 3,966,890 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 96.47 |
| Mean single sequence MFE | -41.32 |
| Consensus MFE | -37.38 |
| Energy contribution | -38.02 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.07 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.16 |
| SVM RNA-class probability | 0.999818 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3966792 98 + 23771897 CAAGAGGCAAGAGCCGCGAGCCCCAAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCCUCC--- ...(((((....((((.((.(((((.((((.....(..((....))..).....(((.((((......))))))).))))))))).))))))))))).--- ( -42.50) >DroSec_CAF1 39141 98 + 1 CAAGAGGCAAGAGCCGCGAGCCCCGAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCCUCC--- ...(((((....((((.((.((((.(((((.....(..((....))..).....(((.((((......))))))).))))).))))))))))))))).--- ( -42.60) >DroSim_CAF1 42634 98 + 1 CAAGAGGCAAGAGCCGCGAGCCCCGAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCCUCC--- ...(((((....((((.((.((((.(((((.....(..((....))..).....(((.((((......))))))).))))).))))))))))))))).--- ( -42.60) >DroEre_CAF1 40212 98 + 1 CAAGAGCCAAGAGCCGCGAGCCCCCAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCCCCC--- ............((((.((.((((((((((.....(..((....))..).....(((.((((......))))))).))))))))))))))))......--- ( -39.40) >DroYak_CAF1 42279 101 + 1 CAAGAGCCAAGAGCCGCGAGCCCCGAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCUCCCUCC ...(((....(((((.((..((((.(((((.....(..((....))..).....(((.((((......))))))).))))).))))..))).)))).))). ( -39.50) >consensus CAAGAGGCAAGAGCCGCGAGCCCCGAGCCGAAAAAGUUCCAAAAGGGCCAAAGAGCCAUGAGCCAAUUCUCAGGUAUGGCUGGGGGUCCGGUGCCUCC___ ...(((((....((((.((.((((.(((((.....(..((....))..).....(((.((((......))))))).))))))))).))))))))))).... (-37.38 = -38.02 + 0.64)

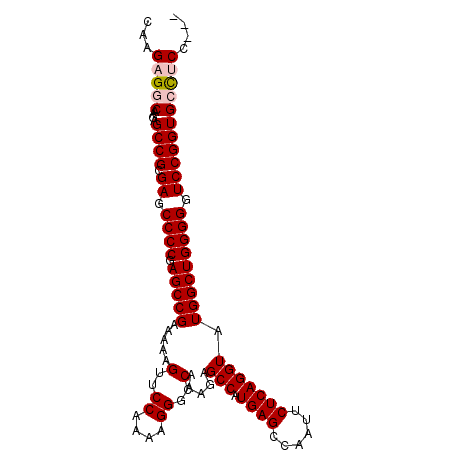
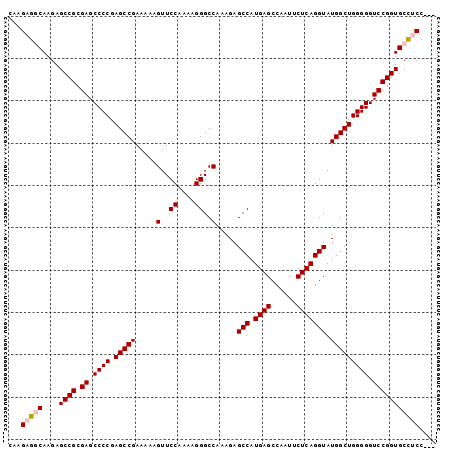
| Location | 3,966,792 – 3,966,890 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 96.47 |
| Mean single sequence MFE | -39.40 |
| Consensus MFE | -34.84 |
| Energy contribution | -35.20 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969315 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3966792 98 - 23771897 ---GGAGGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUUGGGGCUCGCGGCUCUUGCCUCUUG ---((((((((((((((((.((((.....((((......)))).((((....))))................)))).)))).)).)))....))))))).. ( -38.60) >DroSec_CAF1 39141 98 - 1 ---GGAGGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUCGGGGCUCGCGGCUCUUGCCUCUUG ---((((((((((((((((.((((.....((((......)))).((((....))))................)))).)))).)).)))....))))))).. ( -38.60) >DroSim_CAF1 42634 98 - 1 ---GGAGGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUCGGGGCUCGCGGCUCUUGCCUCUUG ---((((((((((((((((.((((.....((((......)))).((((....))))................)))).)))).)).)))....))))))).. ( -38.60) >DroEre_CAF1 40212 98 - 1 ---GGGGGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUGGGGGCUCGCGGCUCUUGGCUCUUG ---((((((..((..(((((((((.....((((......)))).((((....))))................)))))))))..))..))))))........ ( -42.50) >DroYak_CAF1 42279 101 - 1 GGAGGGAGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUCGGGGCUCGCGGCUCUUGGCUCUUG (((((((((..((..((((.((((.....((((......)))).((((....))))................)))).))))..))..)))))...)))).. ( -38.70) >consensus ___GGAGGCACCGGACCCCCAGCCAUACCUGAGAAUUGGCUCAUGGCUCUUUGGCCCUUUUGGAACUUUUUCGGCUCGGGGCUCGCGGCUCUUGCCUCUUG ...((((((((((((((((.((((.....((((......)))).((((....))))................)))).)))).)).)))....))))))).. (-34.84 = -35.20 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:05:35 2006