
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 420,065 – 420,171 |
| Length | 106 |
| Max. P | 0.574424 |

| Location | 420,065 – 420,171 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 82.36 |
| Mean single sequence MFE | -30.38 |
| Consensus MFE | -20.41 |
| Energy contribution | -19.85 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.574424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
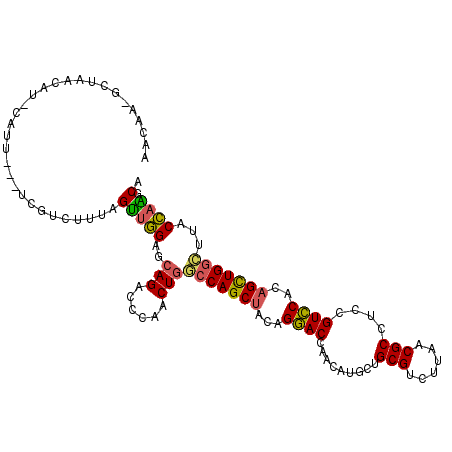
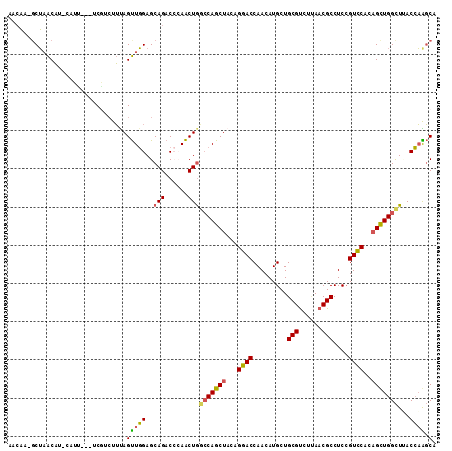
>3L_DroMel_CAF1 420065 106 - 23771897 AACAA-GCUAACAU-CAU----UCGUCUUUAGUUGGAGCAGACGCAACUGACCAGCUAUAGGACCAACAUGCUGCGUCUUAACGCCUCCGUCCACAGUUGGCUUACCAAGCA ...((-((((((..-...----..((((((((((((..(((......))).))))))).))))).........((((....))))...........))))))))........ ( -29.50) >DroVir_CAF1 20394 109 - 1 AACCAUGCGCGCUUUCCUU---CCAACUGCAGCUCGAGCAGACACAGCUAAGCAGCUAUCGCACCAACAUGCUGCGCCUGAACGCCUCGGUGCACAGCUGGUUGACGCGCCA ......(((((.((..(..---....((((.......))))...(((((..(((((..............)))))((((((.....)))).))..))))))..))))))).. ( -34.64) >DroSec_CAF1 1417 107 - 1 AAAAA-UCUAACAU-AAUU---UCGUCUUUAGUUGGAGCAGACCGAACUGGCCAGCUACAGGACCAACAUGCUGCGUCUUAACGCCUCCGUCCACUGUUGGCUUACCAAGCA .....-........-....---..((((((((((((..(((......))).))))))).))))).....((((((((....)))).............(((....))))))) ( -24.60) >DroSim_CAF1 1417 107 - 1 AACAA-GCUAACAU-AAUU---UCGUCUUUAGUUGGAGCAGACCGAACUGGCCAGCUACAGGACCAACAUGCUGCGUCUUAACGCCUCCGUCCACAGCUGGCUUACCAAGCA .....-(((.....-....---.......(((((((......)).)))))(((((((...((((.........((((....))))....))))..)))))))......))). ( -27.12) >DroEre_CAF1 1431 109 - 1 AACAA-GCUAACAU-CAUG-CCGCCCCUUCAGUUGGAGCAGACCCAACUGGCCAGCUACAGGACCAACAUGCUGCGUCUCAACGCCUCCGUCCACAGCUGGCUCACCAGGCA .....-........-..((-((.......(((((((.......)))))))(((((((...((((.........((((....))))....))))..)))))))......)))) ( -35.42) >DroYak_CAF1 1426 110 - 1 AACAA-GCUAACAU-CUUUAUCCCGAAUUUAGUUGGAGCAGACCCAACUGGCCAGCUACAGGACCAACAUGCUGCGUCUCAACGCCUCCGUCCACAGCUGGCUGACCAGGCA .....-(((.....-...............((((((.......))))))((((((((...((((.........((((....))))....))))..)))))))).....))). ( -31.02) >consensus AACAA_GCUAACAU_CAUU___UCGUCUUUAGUUGGAGCAGACCCAACUGGCCAGCUACAGGACCAACAUGCUGCGUCUUAACGCCUCCGUCCACAGCUGGCUUACCAAGCA ...............................(((((..(((......)))(((((((...((((.........(((......)))....))))..)))))))...)))).). (-20.41 = -19.85 + -0.55)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:33:30 2006