
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,867,651 – 3,867,742 |
| Length | 91 |
| Max. P | 0.646064 |

| Location | 3,867,651 – 3,867,742 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 75.96 |
| Mean single sequence MFE | -15.85 |
| Consensus MFE | -8.65 |
| Energy contribution | -8.65 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.646064 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
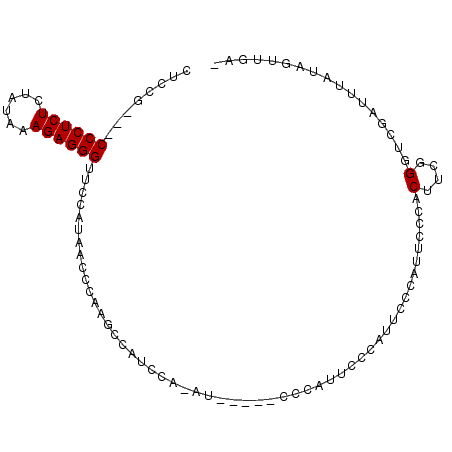
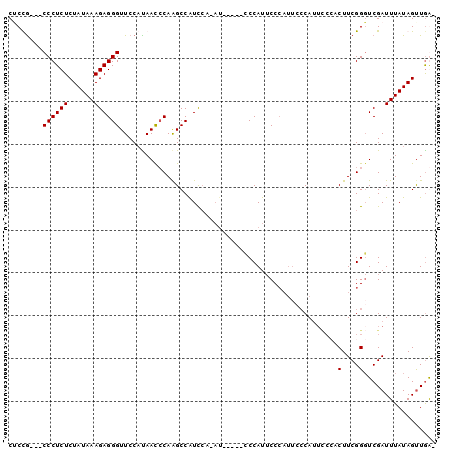
>3L_DroMel_CAF1 3867651 91 + 23771897 CUCUA---CCCUCUCUAUAAAGAGGGUUCCAUAACCCAAGCCAUCCA-AU-----CCCAUUCCCAUUCCCAUUCCCACUUCGGGUCGAUUUAUAGCUGGA ....(---((((((......)))))))........(((.((.....(-((-----(.....(((.................)))..))))....))))). ( -17.33) >DroSec_CAF1 66759 84 + 1 CUCUG---CCCUCUCUAUAAAGAGGGUUCCAUAAUCCAGGCCGUCCA-AU-----CCCAUUCC------CAUUCCCACUUCGGGUCGAUUUAUAGUUGG- ....(---((((((......))))))).................(((-((-----........------....(((.....)))..........)))))- ( -16.25) >DroEre_CAF1 69062 79 + 1 CACCG---CCCUCUCUAUAAAGAGGGUUCCCCAAC-CAAGCCAUU----------UCCAUUCCCAUUCCCA------CUUCGGGUCGAUUUAUAGUUGA- ....(---((((((......)))))))........-.........----------............(((.------....)))(((((.....)))))- ( -14.80) >DroYak_CAF1 69208 99 + 1 CGCCGCUGCCCUCUCUAUAAAGAGGGUUCCAUAACCCAAGCCAUCCACAUCCACAUCCAUACCCAUUCCCAUUUCACCUUCGGCUCGAUUUAUAGUUAA- ....(..(((((((......)))))))..)((((..(.((((.......................................)))).)..))))......- ( -15.03) >consensus CUCCG___CCCUCUCUAUAAAGAGGGUUCCAUAACCCAAGCCAUCCA_AU_____CCCAUUCCCAUUCCCAUUCCCACUUCGGGUCGAUUUAUAGUUGA_ ........((((((......)))))).......................................................................... ( -8.65 = -8.65 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:05:03 2006