
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,858,674 – 3,858,778 |
| Length | 104 |
| Max. P | 0.805092 |

| Location | 3,858,674 – 3,858,778 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 68.83 |
| Mean single sequence MFE | -20.79 |
| Consensus MFE | -12.16 |
| Energy contribution | -11.54 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.805092 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
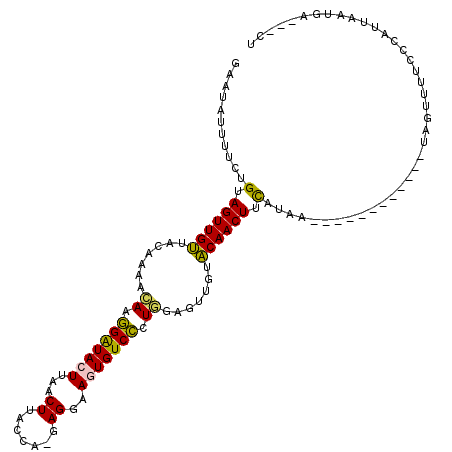
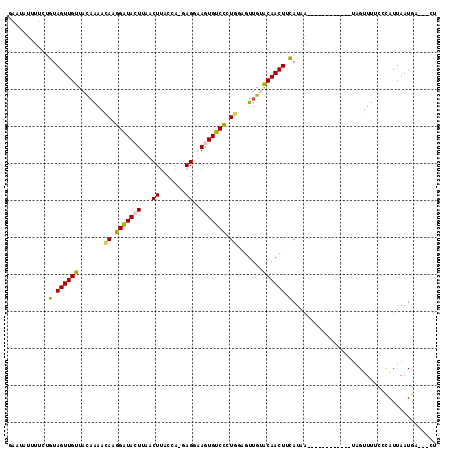
>3L_DroMel_CAF1 3858674 104 - 23771897 GAAUAUUUACCGUAGUUGCUACAGAAUAAGGAUACUUAACUUACCAUUAGUUAGUGUCCCUAGAAUUGUGCAACUUCAUAA------------UAGUUUUCCCAUUAAUGAAGCUU ...........(.((((((.((((..((.(((((((.((((.......))))))))))).))...)))))))))).)....------------.((((((.........)))))). ( -23.20) >DroSec_CAF1 57902 100 - 1 GAAUAUUUUCUGUAGUUGUUAUAAAACAUGGAUAAUUAACUAACUA-GAGGAAGUGUCCCUGGAGUUGUACAACUUCAUAA------------UAGUUUUCCCAUUAAUGA---CU ..(((((((((.(((((((((...............)))).)))))-.)))))))))...(((((((....)))))))...------------..................---.. ( -19.76) >DroSim_CAF1 58517 100 - 1 CAAUAUUUUCUGUAGUUGUUAUAAAACAUGGGUACUUAACUUACAA-GAGAAAGUGUCCCUGGAGUUGCACAACUUCAUAA------------UAGUUUUCCCAUUAAUGA---CU .............(((..(((.(((((..((((((((..(((....-))).))))).)))(((((((....)))))))...------------..))))).....)))..)---)) ( -19.90) >DroYak_CAF1 60209 102 - 1 ----------AGUAGUUGUUUCUAACAAAAGAUACUUAACUUGCCA-UAGGUAAUGUCUAUGGGGUUGUACAACUCUAUAACCAAAAGAGUAAAAGAUUAUUUAUUAAUGA---CG ----------....((..(((((......)))(((((......(((-((((.....)))))))(((((((......)))))))....)))))..............))..)---). ( -20.30) >consensus GAAUAUUUUCUGUAGUUGUUACAAAACAAGGAUACUUAACUUACCA_GAGGAAGUGUCCCUGGAGUUGUACAACUUCAUAA____________UAGUUUUCCCAUUAAUGA___CU ...........(.((((((.......((.(((((((...((.......))..))))))).)).......)))))).)....................................... (-12.16 = -11.54 + -0.62)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:53 2006