
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,782,450 – 3,782,558 |
| Length | 108 |
| Max. P | 0.929521 |

| Location | 3,782,450 – 3,782,558 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 91.03 |
| Mean single sequence MFE | -24.55 |
| Consensus MFE | -21.84 |
| Energy contribution | -21.90 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929521 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
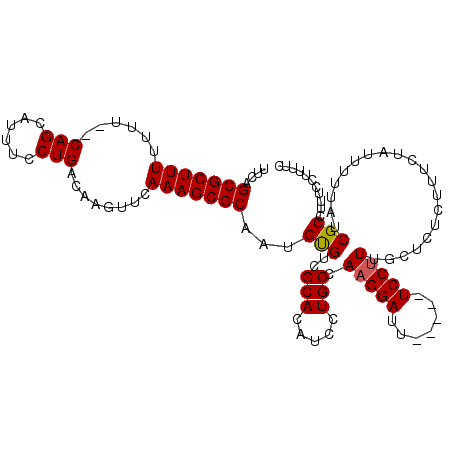
>3L_DroMel_CAF1 3782450 108 + 23771897 GAAAAGGAAAGGCGAUCAAAAUAGAAAGAGAGCAAAGGA-----AAUCCUUGGCAGGAUGUGCGACACAUUGCGCUUUGAACUUGUCAGGAAAUGCUG--AAAAAAAGCGCUGAA ...(((..((((((.................((.((((.-----...)))).))..((((((...)))))).))))))...))).((((....((((.--......)))))))). ( -24.90) >DroSec_CAF1 180052 107 + 1 GAAAAGGAAAGGCGAUAA-AAUAGAAAGAGAGCAAAGGA-----AAUCCUUGGCAGGAUGUGCGACGCAUUGCGCUUUGAACUUGUCAGGAAAUGCUG--AAUAAAAGCGCUGUA ((.(((..((((((....-............((.((((.-----...)))).))..((((((...)))))).))))))...))).)).......(((.--......)))...... ( -23.70) >DroSim_CAF1 174650 108 + 1 GAAAAGGAAAGGCGAUAAAAAUAGAAAGAGAGCAAAGGA-----AAUCCUUGGCAGGAUGUGCGACACAUUGCGCUUUGAACUUGUCAGGAAAUGCUG--AAAAAAAGCGCUGAA ...(((..((((((.................((.((((.-----...)))).))..((((((...)))))).))))))...))).((((....((((.--......)))))))). ( -24.90) >DroYak_CAF1 165167 115 + 1 CAAAAGGGAGAGCGAUAAAAAUAGAAAGAGAGGAAGGGAAGGGAAAUCCUUGGCAGGAUGUGCGACACAUUGCGCUUUGAACUUGCCAGGAAAUGCUGCGAAAAAAAGCGCCGAA .....((...((((....................(((((.......)))))((((((..((((((....))))))......))))))......))))((........)).))... ( -24.70) >consensus GAAAAGGAAAGGCGAUAAAAAUAGAAAGAGAGCAAAGGA_____AAUCCUUGGCAGGAUGUGCGACACAUUGCGCUUUGAACUUGUCAGGAAAUGCUG__AAAAAAAGCGCUGAA ..........(((.................................((((.((((((..((((((....))))))......))))))))))...(((.........))))))... (-21.84 = -21.90 + 0.06)



| Location | 3,782,450 – 3,782,558 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 91.03 |
| Mean single sequence MFE | -18.62 |
| Consensus MFE | -14.63 |
| Energy contribution | -14.69 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3782450 108 - 23771897 UUCAGCGCUUUUUUU--CAGCAUUUCCUGACAAGUUCAAAGCGCAAUGUGUCGCACAUCCUGCCAAGGAUU-----UCCUUUGCUCUCUUUCUAUUUUGAUCGCCUUUCCUUUUC .((((((((((((.(--(((......)))).))....))))))).(((((...)))))...((.((((...-----.)))).)).............)))............... ( -19.80) >DroSec_CAF1 180052 107 - 1 UACAGCGCUUUUAUU--CAGCAUUUCCUGACAAGUUCAAAGCGCAAUGCGUCGCACAUCCUGCCAAGGAUU-----UCCUUUGCUCUCUUUCUAUU-UUAUCGCCUUUCCUUUUC ....(((((((...(--(((......)))).......)))))))...(((......(((((....))))).-----.....)))............-.................. ( -18.30) >DroSim_CAF1 174650 108 - 1 UUCAGCGCUUUUUUU--CAGCAUUUCCUGACAAGUUCAAAGCGCAAUGUGUCGCACAUCCUGCCAAGGAUU-----UCCUUUGCUCUCUUUCUAUUUUUAUCGCCUUUCCUUUUC ....(((((((((.(--(((......)))).))....))))))).(((((...)))))...((.((((...-----.)))).))............................... ( -19.50) >DroYak_CAF1 165167 115 - 1 UUCGGCGCUUUUUUUCGCAGCAUUUCCUGGCAAGUUCAAAGCGCAAUGUGUCGCACAUCCUGCCAAGGAUUUCCCUUCCCUUCCUCUCUUUCUAUUUUUAUCGCUCUCCCUUUUG ...(((((........))......((((((((.(......((((.....).)))....).)))).)))).................................))).......... ( -16.90) >consensus UUCAGCGCUUUUUUU__CAGCAUUUCCUGACAAGUUCAAAGCGCAAUGUGUCGCACAUCCUGCCAAGGAUU_____UCCUUUGCUCUCUUUCUAUUUUUAUCGCCUUUCCUUUUC ....(((((((......(((......)))........)))))))...(((..(((.....))).(((((.......)))))....................)))........... (-14.63 = -14.69 + 0.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:20 2006