
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,688,897 – 3,688,987 |
| Length | 90 |
| Max. P | 0.808828 |

| Location | 3,688,897 – 3,688,987 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 95.74 |
| Mean single sequence MFE | -20.13 |
| Consensus MFE | -15.64 |
| Energy contribution | -16.20 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.556493 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3688897 90 + 23771897 GUUUAUCAGUCGCUGCCGGGCUGGGAAAUAUUCCC-CUU-UUGGAUUUU-CCAGCAUUUUCCUCCCUGAUUUUCCCCUUCCACUGUCAGCAAA ...........((((.((((((((((((...(((.-...-..)))))))-)))))....((......)).............))).))))... ( -22.10) >DroSec_CAF1 84275 90 + 1 GUUUAUCAGUCGCUGCCGGGCUGGGAAGUAUUCCC-CUU-UUGGAUUUU-CCAGCAUUUUCCUCCCUGAUUUUCCCCUUCCACUGUCAACAAA ....(((((..........(((((((((...(((.-...-..)))))))-)))))..........)))))....................... ( -17.45) >DroSim_CAF1 79257 90 + 1 GUUUAUCAGUCGCUGCCGGGCUGGGAAGUAUUCCC-CUU-UUGGAUUUU-CCAGCAUUUUCCUCCCUGAUUUUCCCCUUCCACUGUCAGCAAA ...........((((.((((((((((((...(((.-...-..)))))))-)))))....((......)).............))).))))... ( -21.80) >DroEre_CAF1 78838 91 + 1 GUUUAUCAGUCGCUGCCGGGCUGGGAAAUAUUCCCCCUU-UUGGUUUUU-CCAGCAUUUUUCUCCCUGAUUUUCCCCUUCCACUGUCAGCAAA ...........((((.((((((((((((....((.....-..)).))))-))))).....((.....)).............))).))))... ( -21.80) >DroYak_CAF1 66347 93 + 1 GUUUAUCAGUCGCCGCCGGGCUGGGAAAUAUUCCCACUUUUUGGUUUUUACCAGCAUUUUCCUCCCUGAUUUUCCCCUUCCACUGUCAGCAAA (((...((((.(((....)))(((((.....)))))....(((((....)))))...........................))))..)))... ( -17.50) >consensus GUUUAUCAGUCGCUGCCGGGCUGGGAAAUAUUCCC_CUU_UUGGAUUUU_CCAGCAUUUUCCUCCCUGAUUUUCCCCUUCCACUGUCAGCAAA ...........((((.((((((((.(((...(((........))).))).)))))............((.........))..))).))))... (-15.64 = -16.20 + 0.56)



| Location | 3,688,897 – 3,688,987 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 95.74 |
| Mean single sequence MFE | -22.38 |
| Consensus MFE | -19.54 |
| Energy contribution | -19.22 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.808828 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3688897 90 - 23771897 UUUGCUGACAGUGGAAGGGGAAAAUCAGGGAGGAAAAUGCUGG-AAAAUCCAA-AAG-GGGAAUAUUUCCCAGCCCGGCAGCGACUGAUAAAC .((((((...(.((...((((((.((((.(.......).))))-....(((..-...-.)))...))))))..)))..))))))......... ( -23.40) >DroSec_CAF1 84275 90 - 1 UUUGUUGACAGUGGAAGGGGAAAAUCAGGGAGGAAAAUGCUGG-AAAAUCCAA-AAG-GGGAAUACUUCCCAGCCCGGCAGCGACUGAUAAAC .((((((...(.((...(((((..((((.(.......).))))-....(((..-...-.)))....)))))..)))..))))))......... ( -19.90) >DroSim_CAF1 79257 90 - 1 UUUGCUGACAGUGGAAGGGGAAAAUCAGGGAGGAAAAUGCUGG-AAAAUCCAA-AAG-GGGAAUACUUCCCAGCCCGGCAGCGACUGAUAAAC .((((((...(.((...(((((..((((.(.......).))))-....(((..-...-.)))....)))))..)))..))))))......... ( -22.30) >DroEre_CAF1 78838 91 - 1 UUUGCUGACAGUGGAAGGGGAAAAUCAGGGAGAAAAAUGCUGG-AAAAACCAA-AAGGGGGAAUAUUUCCCAGCCCGGCAGCGACUGAUAAAC ((..(.....)..))........(((((...(.....((((((-.....((..-..))((((.....))))...)))))).)..))))).... ( -22.70) >DroYak_CAF1 66347 93 - 1 UUUGCUGACAGUGGAAGGGGAAAAUCAGGGAGGAAAAUGCUGGUAAAAACCAAAAAGUGGGAAUAUUUCCCAGCCCGGCGGCGACUGAUAAAC .((((((...(.((...(((((((((((.(.......).))))).....(((.....))).....))))))..)))..))))))......... ( -23.60) >consensus UUUGCUGACAGUGGAAGGGGAAAAUCAGGGAGGAAAAUGCUGG_AAAAUCCAA_AAG_GGGAAUAUUUCCCAGCCCGGCAGCGACUGAUAAAC .((((((...(.((...(((((..((((.(.......).))))......((........)).....)))))..)))..))))))......... (-19.54 = -19.22 + -0.32)


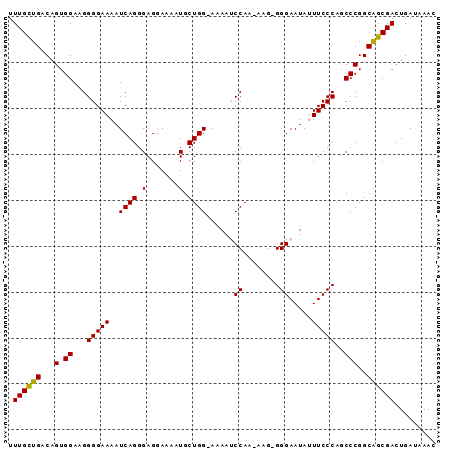
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:03:36 2006