
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,468,928 – 3,469,021 |
| Length | 93 |
| Max. P | 0.741841 |

| Location | 3,468,928 – 3,469,021 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 83.33 |
| Mean single sequence MFE | -18.95 |
| Consensus MFE | -15.10 |
| Energy contribution | -16.73 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.741841 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3468928 93 + 23771897 GGUAUAUAACCUUACACCCCACCCCAAACCACUGAUUAAUACCUUAUCUGCAGCAGAUGCUGUACGUUAUGGGUUAAGUCGAAUCCCCUUACC (((.....))).....................(((((...((((.((.((((((....)))))).))...))))..)))))............ ( -16.70) >DroSec_CAF1 15977 87 + 1 GGUAUAUAACCUUGCACGACACCUUAACCCACUGAUUAAUACCUUAUCUGCAGCAGAUGCU------UAUGGGUUAAGUCGAAUCCCCUUCCC (((.....))).....((((...((((((((..(((.........))).(((.....))).------..))))))))))))............ ( -17.50) >DroSim_CAF1 16344 93 + 1 GGUAUAUAACCUUGCACGCCACCUAAACCCACUGAUUAAUACCUUAUCUGCAGCAGAUGCUGUACGUUAUGGGUUAAGUCGAAUCCCCUUCCC (((.....)))...........((.((((((..(((.........)))((((((....)))))).....)))))).))............... ( -16.90) >DroYak_CAF1 16897 82 + 1 GGUAUCUAACCUUUCACCCUGGCUUAACCCACUGAUUAAUACCUUAUCUGCAGCAGAUGCUGCACGUUAUGGGUUAAGUU-----------CC (((.....))).........(((((((((((..(((.........)))((((((....)))))).....)))))))))))-----------.. ( -24.70) >consensus GGUAUAUAACCUUGCACCCCACCUUAACCCACUGAUUAAUACCUUAUCUGCAGCAGAUGCUGUACGUUAUGGGUUAAGUCGAAUCCCCUUCCC (((.....)))...........(((((((((..(((.........)))((((((....)))))).....)))))))))............... (-15.10 = -16.73 + 1.62)


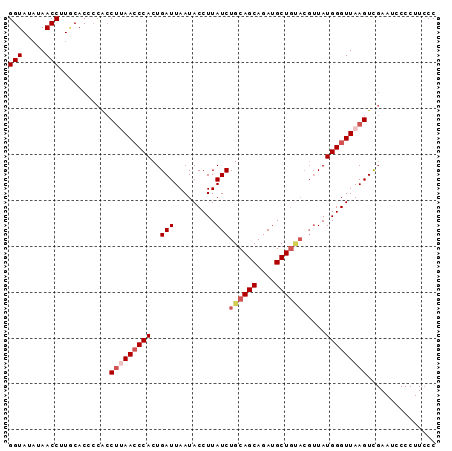
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:33 2006