
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,466,549 – 3,466,708 |
| Length | 159 |
| Max. P | 0.966725 |

| Location | 3,466,549 – 3,466,668 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -22.68 |
| Consensus MFE | -17.86 |
| Energy contribution | -19.36 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.902531 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
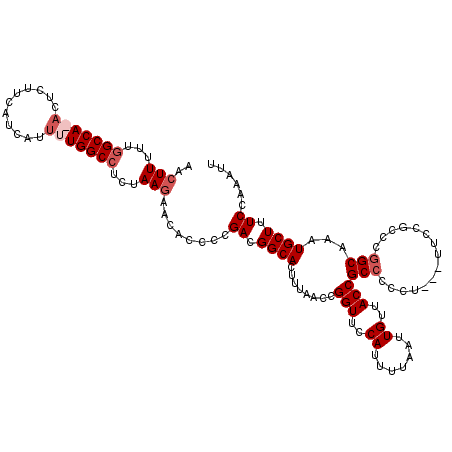
>3L_DroMel_CAF1 3466549 119 + 23771897 AACUUUUUGGCCA-ACUCUUCAUCAUUUUGGCCUCUAAGAACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCACCUUCCUUCCGCCCGGCAAAUGCUUUCCAAAUU ..(((...(((((-(............))))))...)))........((.((((........(((..((.......))..)))(((...............)))...)))).))...... ( -22.96) >DroSec_CAF1 13716 116 + 1 AACUUUUUGGCCAAACUCUUCAUCAUUUUGGCCUCUAAGAAUACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCU---UUCCGCCC-GCAAAUGCUUUCCAAAUU ..(((...(((((((...........)))))))...)))...........(((........((((..((.......))..))))......---....))).-((....)).......... ( -20.77) >DroSim_CAF1 13839 117 + 1 AACUUUUUGGCCAAACUCUUCAUCAUUUUGGCCUCUAAGAACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCA---UUCUGCCCGGCAAAUGCUUUCCAAAUU ..(((...(((((((...........)))))))...)))......(((..((((.......((((..((.......))..))))......---...)))))))................. ( -23.69) >DroYak_CAF1 14292 115 + 1 AACUUUUGGGCCA-ACUCUUCAUCAACUUGGCAUCUAAAAACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCUUC----UUCCGUUCGGCAAAUGCUUUCCAAAUU ....(((((((((-(............))))).............(((((((.........((((..((.......))..)))).....----..))).))))..........))))).. ( -23.29) >consensus AACUUUUUGGCCA_ACUCUUCAUCAUUUUGGCCUCUAAGAACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCU___UUCCGCCCGGCAAAUGCUUUCCAAAUU ..(((...(((((((...........)))))))...)))........((.((((........(((..((.......))..)))(((...............)))...)))).))...... (-17.86 = -19.36 + 1.50)


| Location | 3,466,588 – 3,466,708 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.80 |
| Mean single sequence MFE | -24.39 |
| Consensus MFE | -23.09 |
| Energy contribution | -23.34 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966725 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
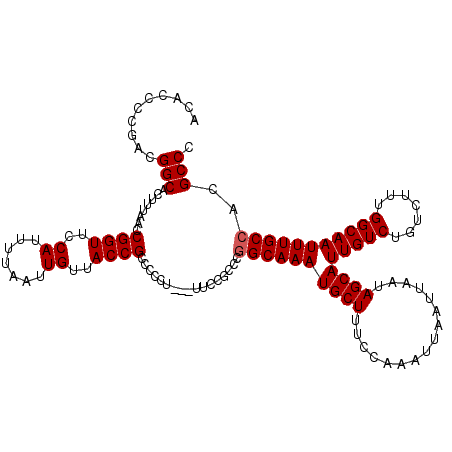
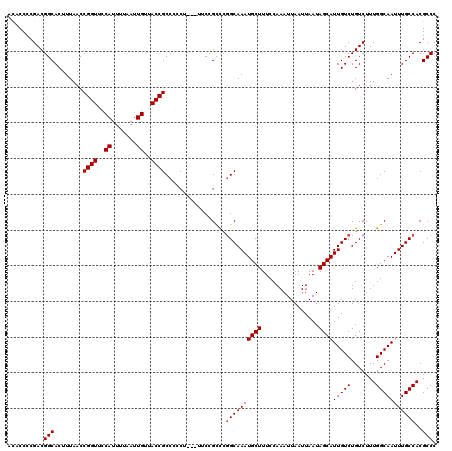
>3L_DroMel_CAF1 3466588 120 + 23771897 ACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCACCUUCCUUCCGCCCGGCAAAUGCUUUCCAAAUUAAUUAAUAGCAUUGUCUGUCUUUGGCAAUUUGCCACGCCC ..........(((........((((..((.......))..)))).................((((((((((................))))(((((.......)))))))))))..))). ( -24.79) >DroSec_CAF1 13756 116 + 1 AUACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCU---UUCCGCCC-GCAAAUGCUUUCCAAAUUAAUUAAUAGCAUUGUCUGUCUUUGGCAAUUUGCCACGCCC .......(((((.((......((((..((.......))..))))......---........-...((((((................)))))))))))))..((((.....))))..... ( -21.99) >DroSim_CAF1 13879 117 + 1 ACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCA---UUCUGCCCGGCAAAUGCUUUCCAAAUUAAUUAAUAGCAUUGUCUGUCUUUGGCAAUUUGCCACGCCC .......(((((((.......((((..((.......))..))))......---...)))).(((((.((((................))))))))).)))..((((.....))))..... ( -25.98) >DroYak_CAF1 14331 116 + 1 ACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCUUC----UUCCGUUCGGCAAAUGCUUUCCAAAUUAAUUAAUAGCAUUGUCUGUCUUUGGCAAUUUGCCACGCCC ..........(((........((((..((.......))..)))).....----........((((((((((................))))(((((.......)))))))))))..))). ( -24.79) >consensus ACACCCCGACGGCACUUUAACCGGUUCCAUUUUAAUUGUUACCGCCCCCU___UUCCGCCCGGCAAAUGCUUUCCAAAUUAAUUAAUAGCAUUGUCUGUCUUUGGCAAUUUGCCACGCCC ..........(((........((((..((.......))..)))).................((((((((((................))))(((((.......)))))))))))..))). (-23.09 = -23.34 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:32 2006