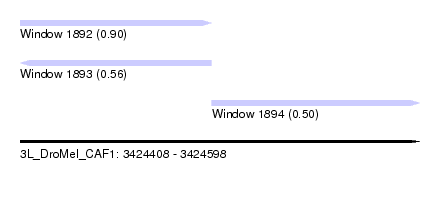

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,424,408 – 3,424,598 |
| Length | 190 |
| Max. P | 0.896700 |

| Location | 3,424,408 – 3,424,499 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 91.23 |
| Mean single sequence MFE | -26.66 |
| Consensus MFE | -22.14 |
| Energy contribution | -23.10 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.896700 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
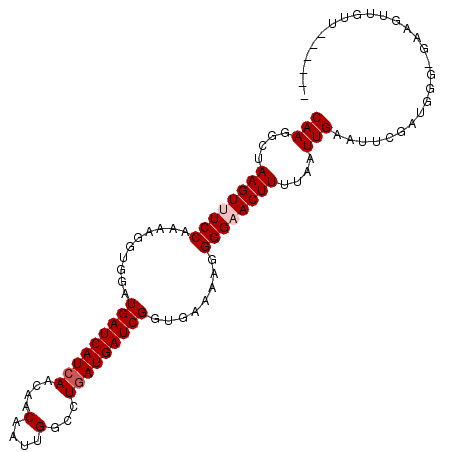
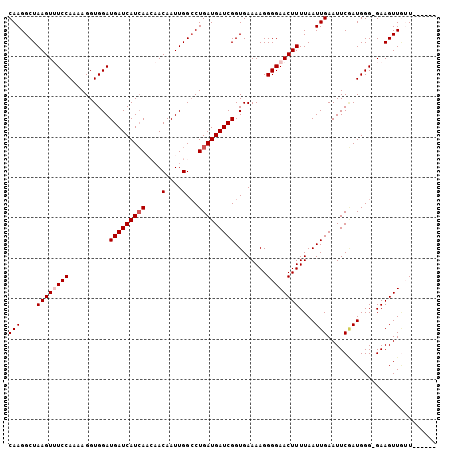
>3L_DroMel_CAF1 3424408 91 + 23771897 GACCGGAGGAUAAACAACGGCCGCGGACUUAAAAAGUAUAC------------AAGAGCAGCGAAGAUUGCCGGUUCGCUGCUUAAUUAGCCGCGGUGACCCU ..(((..(......)..)))(((((((((.....)))....------------..((((((((((.........))))))))))......))))))....... ( -30.20) >DroSec_CAF1 39235 91 + 1 GACCGGAGGAUAAACAACGGCCGCGGACUUAAAAAGUAUAA------------AAGAGCAGCGAAGAUUGCCGGAUCGCUGCUUAAUUAGCCGCGGUGACCCU ..(((..(......)..)))(((((((((.....)))....------------..(((((((((...........)))))))))......))))))....... ( -29.20) >DroSim_CAF1 28086 91 + 1 GACCGGAGGAUAAACAACGGCCGCGGACUUAAAAAGUAUAA------------AAGAGCAGCGAAGAUUGCCGGAUCGCUGCUUAAUUAGCCGCGGUGACCCU ..(((..(......)..)))(((((((((.....)))....------------..(((((((((...........)))))))))......))))))....... ( -29.20) >DroEre_CAF1 39696 91 + 1 GACCGGAGGAUAAACAACGGCUGCAGACUUAAAAAGUAUAA------------AAGAGCAGCGAAGAUUGCCGGAUCGCUGCUUAAUUAGCCACGGUGACCCU .((((..(......)...(((((...(((.....)))....------------..(((((((((...........)))))))))...))))).))))...... ( -25.00) >DroYak_CAF1 38346 103 + 1 GACCGGAGGAUAAACAACGGCUGCAGACUUAAAAUGUAUAAAAAAAAAAAGAAAAGAGCAACGAAGAUUGCCGGUUCGCUGCUUAAUUAGCCACGGUGACCCU .((((..(......)...((((((((((.......)).............(((..(.((((......)))))..))).))))......)))).))))...... ( -19.70) >consensus GACCGGAGGAUAAACAACGGCCGCGGACUUAAAAAGUAUAA____________AAGAGCAGCGAAGAUUGCCGGAUCGCUGCUUAAUUAGCCGCGGUGACCCU ..(((..(......)..)))(((((((((.....)))..................(((((((((...........)))))))))......))))))....... (-22.14 = -23.10 + 0.96)

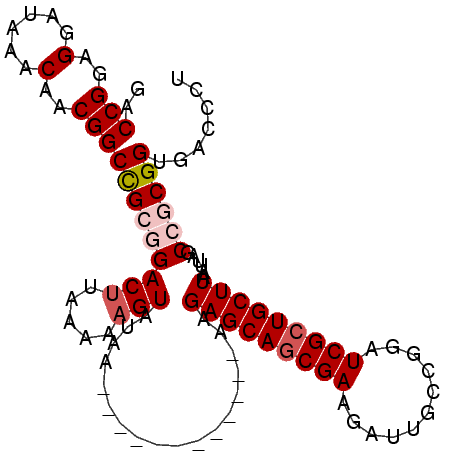
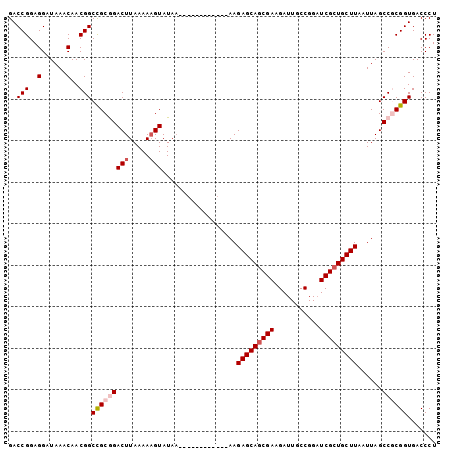
| Location | 3,424,408 – 3,424,499 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 91.23 |
| Mean single sequence MFE | -24.68 |
| Consensus MFE | -23.88 |
| Energy contribution | -23.24 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558718 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3424408 91 - 23771897 AGGGUCACCGCGGCUAAUUAAGCAGCGAACCGGCAAUCUUCGCUGCUCUU------------GUAUACUUUUUAAGUCCGCGGCCGUUGUUUAUCCUCCGGUC (((((.((.((((((.....(((((((((.........)))))))))...------------((..(((.....)))..)))))))).))..)))))...... ( -28.10) >DroSec_CAF1 39235 91 - 1 AGGGUCACCGCGGCUAAUUAAGCAGCGAUCCGGCAAUCUUCGCUGCUCUU------------UUAUACUUUUUAAGUCCGCGGCCGUUGUUUAUCCUCCGGUC (((((.((.((((((.....((((((((...........))))))))...------------....(((.....)))....)))))).))..)))))...... ( -26.00) >DroSim_CAF1 28086 91 - 1 AGGGUCACCGCGGCUAAUUAAGCAGCGAUCCGGCAAUCUUCGCUGCUCUU------------UUAUACUUUUUAAGUCCGCGGCCGUUGUUUAUCCUCCGGUC (((((.((.((((((.....((((((((...........))))))))...------------....(((.....)))....)))))).))..)))))...... ( -26.00) >DroEre_CAF1 39696 91 - 1 AGGGUCACCGUGGCUAAUUAAGCAGCGAUCCGGCAAUCUUCGCUGCUCUU------------UUAUACUUUUUAAGUCUGCAGCCGUUGUUUAUCCUCCGGUC (((((.((..(((((.....((((((((...........))))))))...------------....(((.....)))....)))))..))..)))))...... ( -22.90) >DroYak_CAF1 38346 103 - 1 AGGGUCACCGUGGCUAAUUAAGCAGCGAACCGGCAAUCUUCGUUGCUCUUUUCUUUUUUUUUUUAUACAUUUUAAGUCUGCAGCCGUUGUUUAUCCUCCGGUC (((((.((..(((((.....(((((((((.........)))))))))............................(....))))))..))..)))))...... ( -20.40) >consensus AGGGUCACCGCGGCUAAUUAAGCAGCGAUCCGGCAAUCUUCGCUGCUCUU____________UUAUACUUUUUAAGUCCGCGGCCGUUGUUUAUCCUCCGGUC (((((.((..(((((.....((((((((...........))))))))............................(....))))))..))..)))))...... (-23.88 = -23.24 + -0.64)

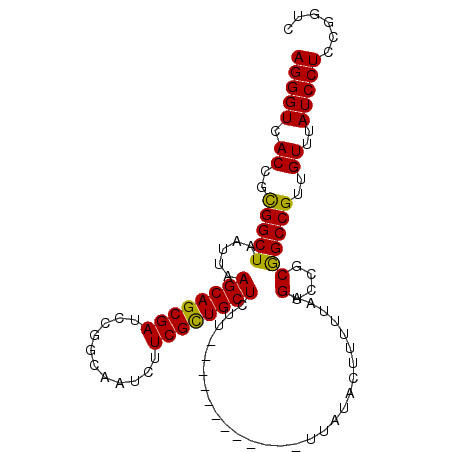
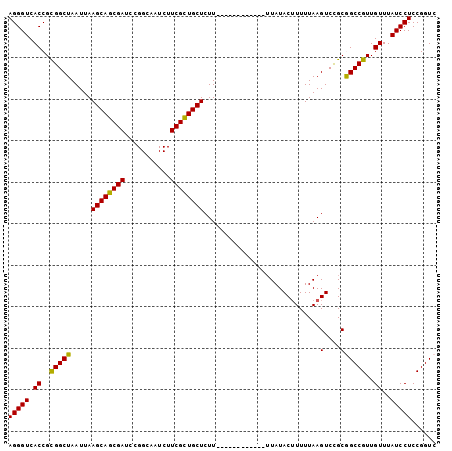
| Location | 3,424,499 – 3,424,598 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 91.45 |
| Mean single sequence MFE | -23.40 |
| Consensus MFE | -18.57 |
| Energy contribution | -19.17 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.79 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3424499 99 + 23771897 CAAGGCUAAGUUUCCAAAAGGUGGAUGAUCAUCAACAACAAUUGGCCUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCGAUGGG-GAAGUUGUU------ (((....((((((((..........(((((((((.((.....))...)))))))))........))))))))....)))....((((...-...))))..------ ( -21.47) >DroSec_CAF1 39326 99 + 1 CAAGGCUAAGUUUCCAAAAGGUGGAUGAUCAUGAACAACAAUUGGCCUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCUAUGGG-GAAGUUGAU------ ......(((.(((((...((((.(((..............))).)))).....(((((..(((((....))))).)))))........))-))).)))..------ ( -19.14) >DroSim_CAF1 28177 99 + 1 CAAGGCUAAGUUUCCAAAAGGUGGAUGAUCAUCAACAACAAUUGGCCUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCGAUGGG-GAAGUUGAU------ (((....((((((((..........(((((((((.((.....))...)))))))))........))))))))....)))...(((((...-...))))).------ ( -23.47) >DroEre_CAF1 39787 97 + 1 CAAGGCUAAGUAUCCAA-AGGUGGAUGAUCAUCA---ACAAUUGGCCUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCGAUGGGGGAAGUUAUUU----- ....(((....(((((.-...)))))((((((((---..........))))))))))).........(((((((.((((....))))...)))))))....----- ( -21.30) >DroYak_CAF1 38449 102 + 1 CAAGGCUAAGUAUCCAAAAGGUGGAUGAUCAUCA---ACAAUUGGACUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCAAUGGG-GAAGUUUUUCUUUUU ....(((....(((((.....)))))((((((((---..........)))))))))))((((((..((((((((.((((....))))..)-)))))))..)))))) ( -31.60) >consensus CAAGGCUAAGUUUCCAAAAGGUGGAUGAUCAUCAACAACAAUUGGCCUGAUGAUCGGUGAAAAGGGGAACUUUUAAUUGAAUUCGAUGGG_GAAGUUGUU______ (((....((((((((..........(((((((((....(....)...)))))))))........))))))))....)))........................... (-18.57 = -19.17 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:05 2006