
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,379,529 – 3,379,727 |
| Length | 198 |
| Max. P | 0.999684 |

| Location | 3,379,529 – 3,379,649 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.79 |
| Mean single sequence MFE | -35.08 |
| Consensus MFE | -32.96 |
| Energy contribution | -32.92 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.94 |
| SVM decision value | 3.88 |
| SVM RNA-class probability | 0.999684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
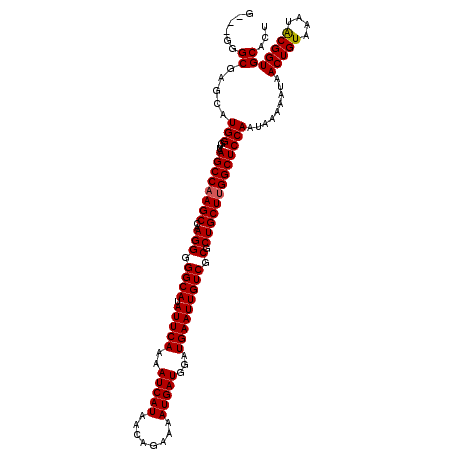
>3L_DroMel_CAF1 3379529 120 + 23771897 GGGGGGGCGAGCAUGGGUUAAGCCAAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAAAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACU ..((((.((((((..((((......))))...(.((((.(((((..(((((........)))))...))))))))).)..)))))).))))..........(((((....)))))..... ( -35.20) >DroSec_CAF1 46627 117 + 1 G---GGGCAAGCAUGGGUUAAGCCCAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAUAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACU (---(((((((((((((.....))))......(.((((.(((((..((((((......))))))...))))))))).)..))))).)))))..........(((((....)))))..... ( -37.00) >DroSim_CAF1 45137 117 + 1 G---GGGCAAGCAUGGGUUAAGCCAAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAAAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACU .---......((.(((....((((((((..(((.((((.(((((..(((((........)))))...))))))))).).))))))))))))).........(((((....)))))))... ( -33.60) >DroEre_CAF1 45967 117 + 1 G---GGGCGACCAUGGGUUAAGCCAAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAAAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUGCGGUGCACU .---((....)).(((....((((((((..(((.((((.(((((..(((((........)))))...))))))))).).))))))))))))).........(((((....)))))..... ( -34.80) >DroYak_CAF1 45804 117 + 1 G---GGGCGACCAUGGGUUAAGCCAAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAAAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUGCGGUGCACU .---((....)).(((....((((((((..(((.((((.(((((..(((((........)))))...))))))))).).))))))))))))).........(((((....)))))..... ( -34.80) >consensus G___GGGCGAGCAUGGGUUAAGCCAAGCCAAGGGGGCAUAUUCAAAAUCAUAACAGAAAAUGAUGGAUGAAUUGUCGCGCUGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACU ......((.....(((....((((((((..(((.((((.(((((..(((((........)))))...))))))))).).))))))))))))).........(((((....)))))))... (-32.96 = -32.92 + -0.04)


| Location | 3,379,609 – 3,379,727 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.65 |
| Mean single sequence MFE | -28.51 |
| Consensus MFE | -24.12 |
| Energy contribution | -24.28 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691866 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3379609 118 + 23771897 UGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACUUUUCCGAAA-GGGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCACGUCGUCGCCCCCCACAAAUCAACAA-AAACAUGCA .((((((((............(((((....)))))...(((..((....-))..)))...))))))......((((((......))))))....................-......)). ( -29.30) >DroSec_CAF1 46704 119 + 1 UGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACUUUUCCGAAA-GGGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCACGUCGUCGCCCCCCACAAAUCAACAAAAAACAUGCA .((((((((............(((((....)))))...(((..((....-))..)))...))))))......((((((......))))))...........................)). ( -29.30) >DroSim_CAF1 45214 118 + 1 UGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACUUUUCCGAAA-GGGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCGCGUCGUCGCCCCCCACAAAUCAACAA-AAACAUGCA .((((((((............(((((....)))))...(((..((....-))..)))...))))))......(((((((....)))))))....................-......)). ( -32.30) >DroEre_CAF1 46044 118 + 1 UGCUUGGCUCCAAUAAAAAUAACUGUAAAUGCGGUGCACUUUUCCGAAA--GGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCACGUCGCCGCCCCCCACAAAUCGACACAAAACAUGCA ((((.((((((..........(((((....)))))........((....--))))).((((............))))))))))).((((...............))))............ ( -25.06) >DroYak_CAF1 45881 120 + 1 UGCUUGGCUCCAAUAAAAAUAACUGUAAAUGCGGUGCACUUUUCCGAAAAGGGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCACGUCGUCGCCCCCCACAAAUCGACACAAAACAUGCA .((((((((............(((((....)))))...(((..((.....))..)))...))))))......((((((......))))))...........................)). ( -26.60) >consensus UGCUUGGCUCCAAUAAAAAUAACUGUAAAUACGGUGCACUUUUCCGAAA_GGGGGAGAUCGGCCAAAUUAAAACGAUGCCAGCACGUCGUCGCCCCCCACAAAUCAACAAAAAACAUGCA .((((((((............(((((....)))))...(((..((.....))..)))...))))))......((((((......))))))...........................)). (-24.12 = -24.28 + 0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:40 2006