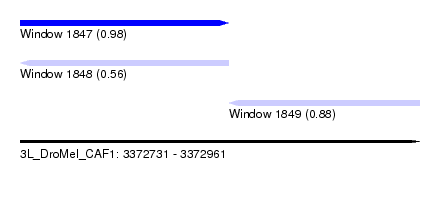

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,372,731 – 3,372,961 |
| Length | 230 |
| Max. P | 0.983950 |

| Location | 3,372,731 – 3,372,851 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.00 |
| Mean single sequence MFE | -33.56 |
| Consensus MFE | -33.34 |
| Energy contribution | -33.74 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.983950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3372731 120 + 23771897 GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUUCGAGGAGAAGGCCUUUGUAUUUGGUUUAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..(((((((..(.((((((((((((((.((.((((.....)))).)).)))).)))))))).)).)..))))))).)))))...)))))................ ( -35.50) >DroSec_CAF1 39864 120 + 1 GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUUCGAGGAGAAGGCCUUUGUAUUUGCUUCAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..((((((.((.((((((.((((((((.((.((((.....)))).)).)))).))))..)).)))).)))))))).)))))...)))))................ ( -30.20) >DroSim_CAF1 37587 120 + 1 GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUUCGAGGAGAAGGCCUUUGUAUUUGGUUUAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..(((((((..(.((((((((((((((.((.((((.....)))).)).)))).)))))))).)).)..))))))).)))))...)))))................ ( -35.50) >DroEre_CAF1 39305 120 + 1 GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUUCGAGGAGAAGGCCUUUGUAUUUGGUUUAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..(((((((..(.((((((((((((((.((.((((.....)))).)).)))).)))))))).)).)..))))))).)))))...)))))................ ( -35.50) >DroYak_CAF1 38602 120 + 1 GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUGCGAGGAGAAGGCCUUUGUAUUUGGUUUAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..(((((((..(.((((((((((((((.((.((.(.....).)).)).)))).)))))))).)).)..))))))).)))))...)))))................ ( -31.10) >consensus GUUAAGCCGCACGCACUAAAACCGAGCCAAAGACCAAGACGACGGACUUCGAGGAGAAGGCCUUUGUAUUUGGUUUAUUUGUCUGGUUUUUUGUGUACAGGUUUUCUUUUAUUCAUUAUU ...(((((..(((((..((((((.((.(((((((((((((((.((.((((.....)))).)).)))).)))))))..)))).)))))))).)))))...)))))................ (-33.34 = -33.74 + 0.40)
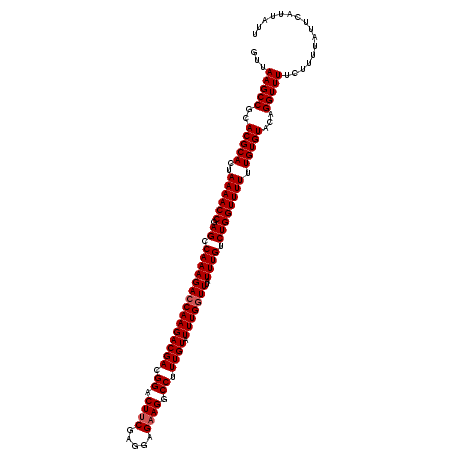


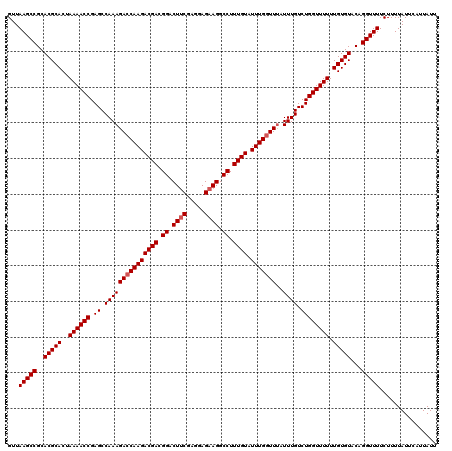
| Location | 3,372,731 – 3,372,851 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.00 |
| Mean single sequence MFE | -26.37 |
| Consensus MFE | -25.34 |
| Energy contribution | -25.74 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562197 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3372731 120 - 23771897 AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUAAACCAAAUACAAAGGCCUUCUCCUCGAAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((((.((((...(((((..((...((.((((.....)))).))...)).))))).)))).)).))))))..)).))))))...... ( -27.50) >DroSec_CAF1 39864 120 - 1 AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUGAAGCAAAUACAAAGGCCUUCUCCUCGAAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((...(....).........((((((((....(..((.....))..)....))))))))....))))))..)).))))))...... ( -25.30) >DroSim_CAF1 37587 120 - 1 AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUAAACCAAAUACAAAGGCCUUCUCCUCGAAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((((.((((...(((((..((...((.((((.....)))).))...)).))))).)))).)).))))))..)).))))))...... ( -27.50) >DroEre_CAF1 39305 120 - 1 AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUAAACCAAAUACAAAGGCCUUCUCCUCGAAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((((.((((...(((((..((...((.((((.....)))).))...)).))))).)))).)).))))))..)).))))))...... ( -27.50) >DroYak_CAF1 38602 120 - 1 AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUAAACCAAAUACAAAGGCCUUCUCCUCGCAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((((................((((((((....(..((.....))..)....)))))))).)).))))))..)).))))))...... ( -24.03) >consensus AAUAAUGAAUAAAAGAAAACCUGUACACAAAAAACCAGACAAAUAAACCAAAUACAAAGGCCUUCUCCUCGAAGUCCGUCGUCUUGGUCUUUGGCUCGGUUUUAGUGCGUGCGGCUUAAC ....................((((((.((.((((((((.((((...(((((..((...((.((((.....)))).))...)).))))).)))).)).))))))..)).))))))...... (-25.34 = -25.74 + 0.40)
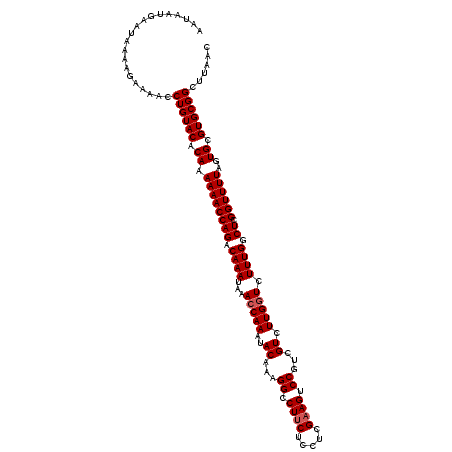


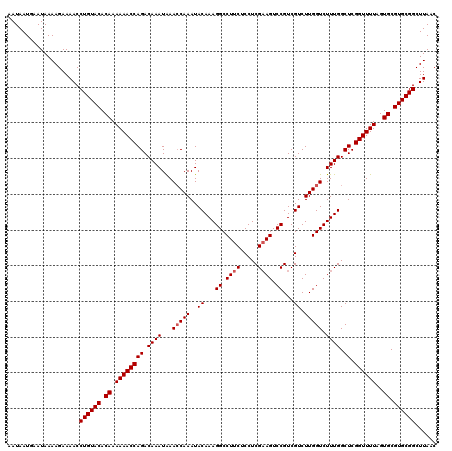
| Location | 3,372,851 – 3,372,961 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 92.63 |
| Mean single sequence MFE | -41.27 |
| Consensus MFE | -38.83 |
| Energy contribution | -38.64 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880504 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 3372851 110 - 23771897 CUCAGCGCAUUUUCGCCCAGUGUAUUCCACAUAAAGCGUAGCACGGUAGCCGAGCUACAGGUUCGAGUGCCGCUGUGUGU------GUGUGUGUGGACGAUGACGCUGGUAAAA-AA ....(((......)))(((((((..((((((((..((...((((((((..(((((.....)))))..))))).)))..))------...)))))))).....))))))).....-.. ( -41.20) >DroSec_CAF1 39984 109 - 1 CUCUGCGCAUUUUCGCCCAGUGUAUUCCACAUAAAGCGUAGCACGGUAGCCGAGCUACAGGUUCGAGUGCCGCUGUGUGU------GUGUGUGUGGACGAUGACGCUGGUAAAA--A ....(((......)))(((((((..((((((((..((...((((((((..(((((.....)))))..))))).)))..))------...)))))))).....))))))).....--. ( -41.00) >DroSim_CAF1 37707 115 - 1 CUGCGCGCAUUUUCGCCCAGUGUAUUCCGCAUAAAGCGUAGCACGGUAGCCGAGCUACAGGUUCGAGUGCCGCUGUGUGUGUCUGUGUGUGUGUGGACGAUGACGCUGGUAAAA--A ....(((......)))(((((((..((((((((..(((..((((((..(((((((.....)))))...))..)))))).....)))...)))))))).....))))))).....--. ( -41.50) >DroYak_CAF1 38722 117 - 1 CUCUGCGCAUUUUCGCCCAGUGUAUUCCACAUAAAGCGUAGCACGGUAGCCGAGCUACAGGUUCGAGUGCGGCUGUGUGUGUGUACUUGUGUGUGGACGAUGACGCUGGUAAAAAAA ....(((......)))(((((((..((((((((....(((((.(((...))).))))).....((((((((((.....)).)))))))))))))))).....)))))))........ ( -41.40) >consensus CUCUGCGCAUUUUCGCCCAGUGUAUUCCACAUAAAGCGUAGCACGGUAGCCGAGCUACAGGUUCGAGUGCCGCUGUGUGU______GUGUGUGUGGACGAUGACGCUGGUAAAA__A ....(((......)))(((((((..((((((((.......((((((..(((((((.....)))))...))..))))))...........)))))))).....)))))))........ (-38.83 = -38.64 + -0.19)

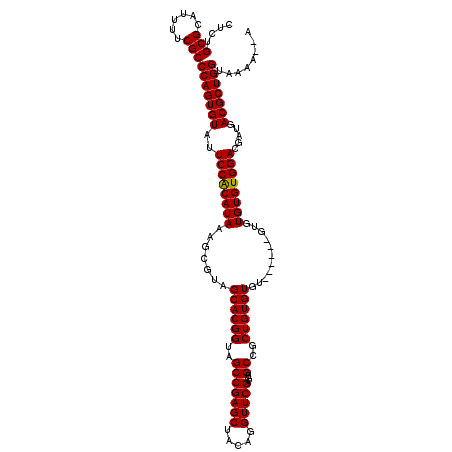
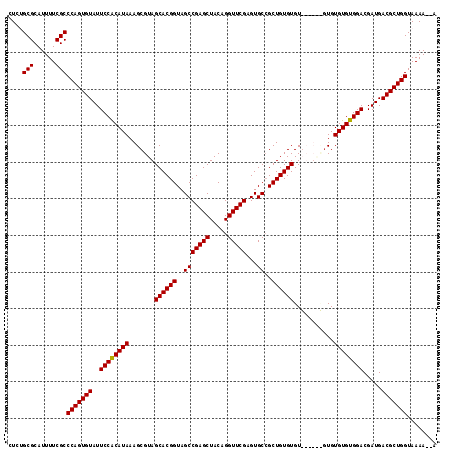
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:23 2006