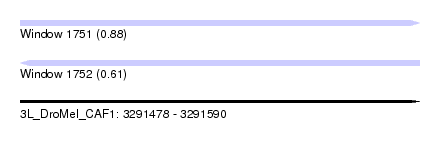

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,291,478 – 3,291,590 |
| Length | 112 |
| Max. P | 0.880066 |

| Location | 3,291,478 – 3,291,590 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.41 |
| Mean single sequence MFE | -43.68 |
| Consensus MFE | -33.00 |
| Energy contribution | -32.64 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
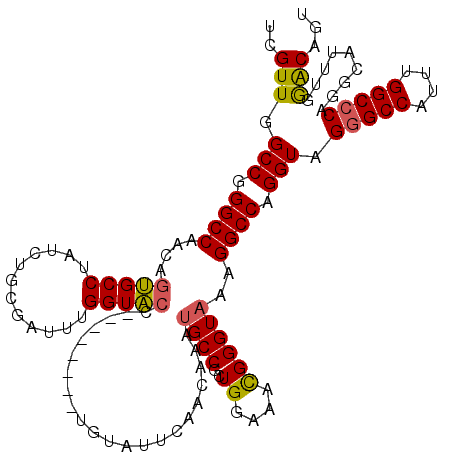
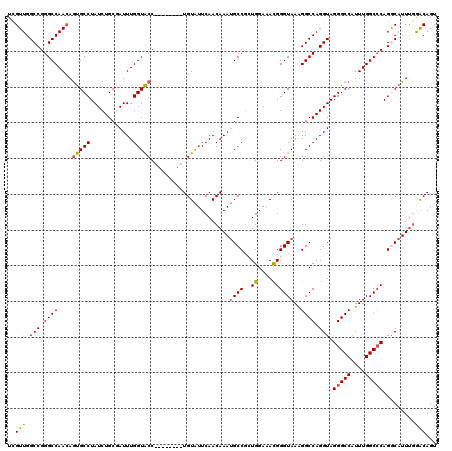
>3L_DroMel_CAF1 3291478 112 + 23771897 ACGUUGGCCGGGCCAACAGUGCCUAUCUGCGAUUUGGUACC--------UGUAUUCAACAAAUGCCGCUGGAAACGGGUAAAGGCCAGGUAGGGCCAUUUGGCACAGGCAUUUGGGCAGU ..((((((...)))))).(((((.(((...)))..)))))(--------(((......((((((((.(((....))).....((((......))))..........)))))))).)))). ( -42.20) >DroSec_CAF1 21237 112 + 1 UCGUUGGCCGGGCCAACAGUGCCUAUCUGCGAUUUGGUACC--------UGUAUUCAACAAAUGCCGCUGGAAAUGGGUAAAGGCCAGGUAGGGCCAUUUGGCCCAGACAUUUGGACAGU ..(((.(((.((((.....(((((((.(.((...(((((..--------(((.....)))..))))).)).).)))))))..)))).))).(((((....))))).........)))... ( -38.20) >DroSim_CAF1 22042 112 + 1 UCGUUGGCCGGGCCAACAGUGCCUAUCUGCGAUUUGGUACC--------UGUAUUCAACAAAUGCCGCUGGAAACGGGUAAAGGCCAGGUAGGGCCAUUUGGCCCAGACAUUUGGACAGU ..(((.(((.((((.((((((((.(((...)))..)))).)--------)))..........((((.(.(....))))))..)))).))).(((((....))))).........)))... ( -39.40) >DroEre_CAF1 23296 112 + 1 UCGUUGGCCGGGCCAACAGUGCCUAUCUGCCAUUUGGUACC--------UGUAUUCAACAAAUGCCGGUGGAAACGGGUAAGGGCCAGGUAGGGCCGUUUGGCCCAGGCAUUCCAGCAGU ..(((((((((((((((.((.((((((((((......((((--------(((.((((.(.......).)))).)))))))..)).)))))))))).).))))))).)))....))))... ( -47.50) >DroYak_CAF1 23961 120 + 1 UUGUUGGCCGGGCCAAAAACGCCUAUCUGCCAUUUGGUGCCAGUACCUGUGUGUUCAACAAAAGCCCGUGGAAACGGGUAAAGGCCAGGUAGGGCCAUUUGGCCCAGGCAUUUGAACAGU (((((.(((((((((((....((((((((((....((((....))))..(((.....)))...((((((....))))))...)).))))))))....)))))))).))).....))))). ( -51.10) >consensus UCGUUGGCCGGGCCAACAGUGCCUAUCUGCGAUUUGGUACC________UGUAUUCAACAAAUGCCGCUGGAAACGGGUAAAGGCCAGGUAGGGCCAUUUGGCCCAGGCAUUUGGACAGU ..(((.(((.((((....(((((............)))))......................((((..((....))))))..)))).))).(((((....))))).........)))... (-33.00 = -32.64 + -0.36)

| Location | 3,291,478 – 3,291,590 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.41 |
| Mean single sequence MFE | -36.82 |
| Consensus MFE | -29.42 |
| Energy contribution | -29.82 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.608175 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
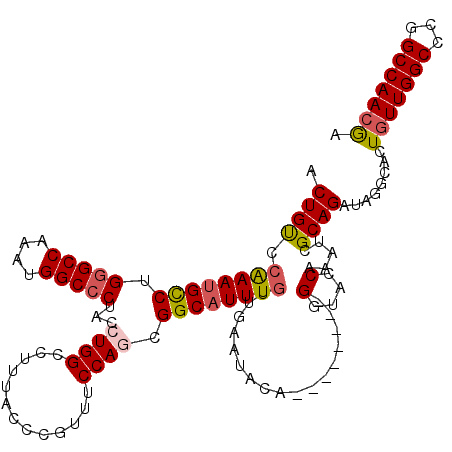
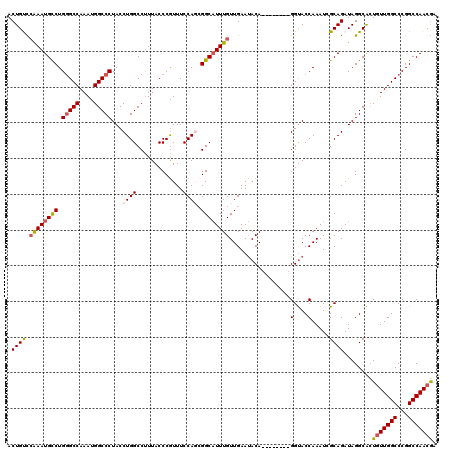
>3L_DroMel_CAF1 3291478 112 - 23771897 ACUGCCCAAAUGCCUGUGCCAAAUGGCCCUACCUGGCCUUUACCCGUUUCCAGCGGCAUUUGUUGAAUACA--------GGUACCAAAUCGCAGAUAGGCACUGUUGGCCCGGCCAACGU .((((.....((((((((.(((((((((......)))).....((((.....)))).))))).....))))--------)))).......)))).........((((((...)))))).. ( -33.60) >DroSec_CAF1 21237 112 - 1 ACUGUCCAAAUGUCUGGGCCAAAUGGCCCUACCUGGCCUUUACCCAUUUCCAGCGGCAUUUGUUGAAUACA--------GGUACCAAAUCGCAGAUAGGCACUGUUGGCCCGGCCAACGA .(((((((((((((.(((((....)))))...((((.............)))).)))))))).........--------(((.....)))...)))))....(((((((...))))))). ( -34.32) >DroSim_CAF1 22042 112 - 1 ACUGUCCAAAUGUCUGGGCCAAAUGGCCCUACCUGGCCUUUACCCGUUUCCAGCGGCAUUUGUUGAAUACA--------GGUACCAAAUCGCAGAUAGGCACUGUUGGCCCGGCCAACGA .(((((.........(((((....)))))((((((........((((.....))))((.....))....))--------))))..........)))))....(((((((...))))))). ( -34.90) >DroEre_CAF1 23296 112 - 1 ACUGCUGGAAUGCCUGGGCCAAACGGCCCUACCUGGCCCUUACCCGUUUCCACCGGCAUUUGUUGAAUACA--------GGUACCAAAUGGCAGAUAGGCACUGUUGGCCCGGCCAACGA ..((((....((((.(((((....)))))((((((........(((.......)))((.....))....))--------))))......))))....)))).(((((((...))))))). ( -38.70) >DroYak_CAF1 23961 120 - 1 ACUGUUCAAAUGCCUGGGCCAAAUGGCCCUACCUGGCCUUUACCCGUUUCCACGGGCUUUUGUUGAACACACAGGUACUGGCACCAAAUGGCAGAUAGGCGUUUUUGGCCCGGCCAACAA ..((((.....(((.((((((((..(((.((((((.......(((((....)))))....(((.....)))))))))(((.((.....)).)))...)))...))))))))))).)))). ( -42.60) >consensus ACUGUCCAAAUGCCUGGGCCAAAUGGCCCUACCUGGCCUUUACCCGUUUCCAGCGGCAUUUGUUGAAUACA________GGUACCAAAUCGCAGAUAGGCACUGUUGGCCCGGCCAACGA .((((.((((((((.(((((....)))))...((((.............)))).)))))))).................(....).....))))........(((((((...))))))). (-29.42 = -29.82 + 0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:59:48 2006