
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 3,066,390 – 3,066,484 |
| Length | 94 |
| Max. P | 0.995453 |

| Location | 3,066,390 – 3,066,484 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -25.25 |
| Consensus MFE | -19.02 |
| Energy contribution | -20.62 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952275 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
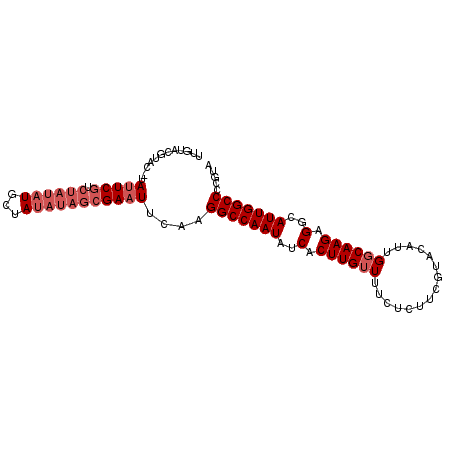
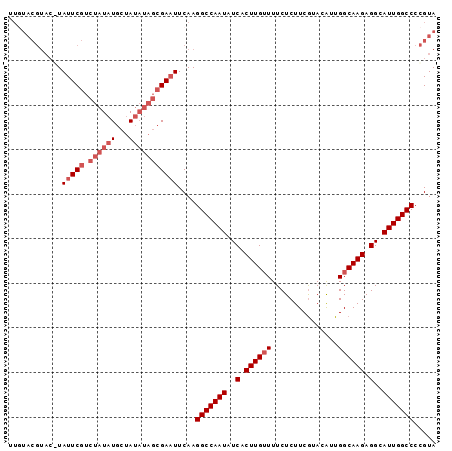
>3L_DroMel_CAF1 3066390 94 + 23771897 UUGUACGUA--UAUUCGUCUAUAUGCUAUAUAGCGAAUUCAAGGCCAAUAUCACUUGUUUUCUCUUCGUACAUUGGCAAGAGGCAUUGGCCCCGUA ...((((..--.(((((.((((((...)))))))))))....(((((((..(.((((((...............)))))).)..))))))).)))) ( -26.06) >DroSec_CAF1 27212 96 + 1 UUGUACGUACGUAUUCGUCUAUAUGCUAUAUAGCGAAUUCAAGGCCAAUAUCACUUGUUUUCUCUUCGUACAUUGGCAAGAGGCAUUGGCCCCGUA ...((((...(.(((((.((((((...))))))))))).)..(((((((..(.((((((...............)))))).)..))))))).)))) ( -27.16) >DroSim_CAF1 27178 96 + 1 UUGUACGUACGUAUUCGUCUAUAUGCUAUAUAGCGAAUUCAAGGCCAAUAUCACUUGUUUUCUCUUCGUACAUUGGCAAGAGGCAUUGGCCCCGUA ...((((...(.(((((.((((((...))))))))))).)..(((((((..(.((((((...............)))))).)..))))))).)))) ( -27.16) >DroYak_CAF1 27417 92 + 1 UUUUACG----UAUUCGUCUAUAUGCUAUAUAGCGAUUUCAAGGCCAAUAUCACUUGUUUUCUCUUCAUACAUUGGCAAGAGGCAUUGGCCCCGUA ...((((----...(((.((((((...)))))))))......(((((((..(.((((((...............)))))).)..))))))).)))) ( -24.56) >DroAna_CAF1 29110 85 + 1 U------UACUUAUUCAAGUAUAUGCGA-----AGAAUUCAAGGCCAAUAUCACUUGAUUUCUCUUCGGACGUUGGCAAGAGGCAUUGGCCCGGUA .------(((((....))))).......-----.........(((((((..(.((((....(((....)).)....)))).)..)))))))..... ( -21.30) >consensus UUGUACGUAC_UAUUCGUCUAUAUGCUAUAUAGCGAAUUCAAGGCCAAUAUCACUUGUUUUCUCUUCGUACAUUGGCAAGAGGCAUUGGCCCCGUA ............(((((.((((((...)))))))))))....(((((((..(.((((((...............)))))).)..)))))))..... (-19.02 = -20.62 + 1.60)


| Location | 3,066,390 – 3,066,484 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -24.47 |
| Consensus MFE | -21.47 |
| Energy contribution | -23.07 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.88 |
| SVM decision value | 2.58 |
| SVM RNA-class probability | 0.995453 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
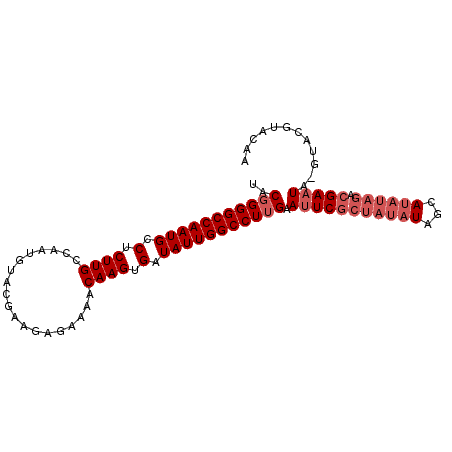
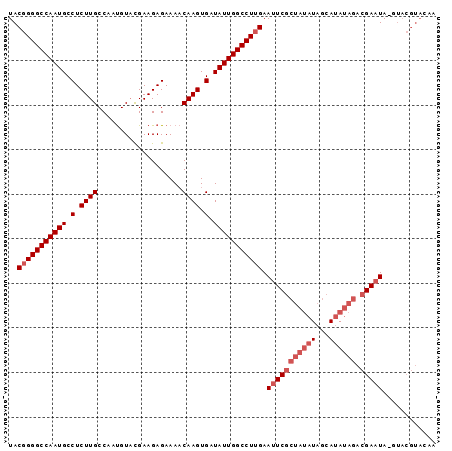
>3L_DroMel_CAF1 3066390 94 - 23771897 UACGGGGCCAAUGCCUCUUGCCAAUGUACGAAGAGAAAACAAGUGAUAUUGGCCUUGAAUUCGCUAUAUAGCAUAUAGACGAAUA--UACGUACAA ..(((((((((((.(.((((...................)))).).))))))))))).(((((((((((...)))))).))))).--......... ( -26.51) >DroSec_CAF1 27212 96 - 1 UACGGGGCCAAUGCCUCUUGCCAAUGUACGAAGAGAAAACAAGUGAUAUUGGCCUUGAAUUCGCUAUAUAGCAUAUAGACGAAUACGUACGUACAA ..(((((((((((.(.((((...................)))).).))))))))))).(((((((((((...)))))).)))))............ ( -26.51) >DroSim_CAF1 27178 96 - 1 UACGGGGCCAAUGCCUCUUGCCAAUGUACGAAGAGAAAACAAGUGAUAUUGGCCUUGAAUUCGCUAUAUAGCAUAUAGACGAAUACGUACGUACAA ..(((((((((((.(.((((...................)))).).))))))))))).(((((((((((...)))))).)))))............ ( -26.51) >DroYak_CAF1 27417 92 - 1 UACGGGGCCAAUGCCUCUUGCCAAUGUAUGAAGAGAAAACAAGUGAUAUUGGCCUUGAAAUCGCUAUAUAGCAUAUAGACGAAUA----CGUAAAA ..(((((((((((.(.((((...................)))).).)))))))))))...(((((((((...)))))).)))...----....... ( -24.61) >DroAna_CAF1 29110 85 - 1 UACCGGGCCAAUGCCUCUUGCCAACGUCCGAAGAGAAAUCAAGUGAUAUUGGCCUUGAAUUCU-----UCGCAUAUACUUGAAUAAGUA------A ..(.(((((((((.(.((((......((......))...)))).).))))))))).)......-----.......(((((....)))))------. ( -18.20) >consensus UACGGGGCCAAUGCCUCUUGCCAAUGUACGAAGAGAAAACAAGUGAUAUUGGCCUUGAAUUCGCUAUAUAGCAUAUAGACGAAUA_GUACGUACAA ..(((((((((((.(.((((...................)))).).))))))))))).(((((((((((...)))))).)))))............ (-21.47 = -23.07 + 1.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:57:23 2006