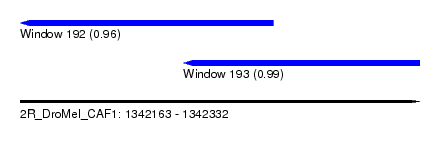
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 1,342,163 – 1,342,332 |
| Length | 169 |
| Max. P | 0.992828 |

| Location | 1,342,163 – 1,342,270 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.71 |
| Mean single sequence MFE | -40.62 |
| Consensus MFE | -37.02 |
| Energy contribution | -38.30 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.957163 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 1342163 107 - 20766785 CGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACGCCAUAAAUUAUGUCGGAGGCUGUCUGUACCAAGUGCACUCGCACAGCCGAACAGGUCAGCCCUUUCUGU----------- .(((..((((((((((.--...))))))))))..))).((.((((...)))).))..((((((((((.(...((((....))))....).))))).)))))........----------- ( -38.70) >DroSec_CAF1 60508 107 - 1 CGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACGCCAUAAAUUAUGUCGGAGGCUGUCUGUACCGGGUGCACUCGCACAGCCGAACAGGUCAGCCCUUUCUGU----------- .(((..((((((((((.--...))))))))))..))).((.((((...)))).))..((((((((((..(((((((....))))..))).))))).)))))........----------- ( -42.40) >DroSim_CAF1 38621 107 - 1 CGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACGCCAUAAAUUAUGUCGGAGGCUGUCUGUACCGGGUGCACUCGCACAGCCGAACAGGUCAGCCCUUUCUGU----------- .(((..((((((((((.--...))))))))))..))).((.((((...)))).))..((((((((((..(((((((....))))..))).))))).)))))........----------- ( -42.40) >DroEre_CAF1 41879 118 - 1 CGCCGGAGACUCGUUUC--GGGAAGCGAGUUUAAGGAACGCCAUAAAUUAUGUCGGAGGCUGUCUGUACCGGGUGCACUCACACAGCCGAGCAGGUCAGCCCUUUCUGUGCCUGUGCCUG .(((((((((((((((.--...)))))))))).(((((((.((((...)))).))..((((((((((..(((.((........)).))).))))).))))).)))))....))).))... ( -39.50) >DroYak_CAF1 29154 109 - 1 CGCCAGAGACUCGUUUCGCGGGAAGCGAGUUUAAGGCACGCAAUAAAUUAUGUCGGAGGCUGUCUGUACCGGGUGCACUUACACAGCCGACCAGGUCAGCCCUUUCUCU----------- .(((..(((((((((((....)))))))))))..)))..............(..((.(((((((((...(((.((........)).)))..)))).)))))))..)...----------- ( -40.10) >consensus CGCCAGAGACUCGUUUC__GGGAAGCGAGUUUAAGGCACGCCAUAAAUUAUGUCGGAGGCUGUCUGUACCGGGUGCACUCGCACAGCCGAACAGGUCAGCCCUUUCUGU___________ .(((..(((((((((((....)))))))))))..))).((.((((...)))).))..((((((((((..(((((((....))))..))).))))).)))))................... (-37.02 = -38.30 + 1.28)

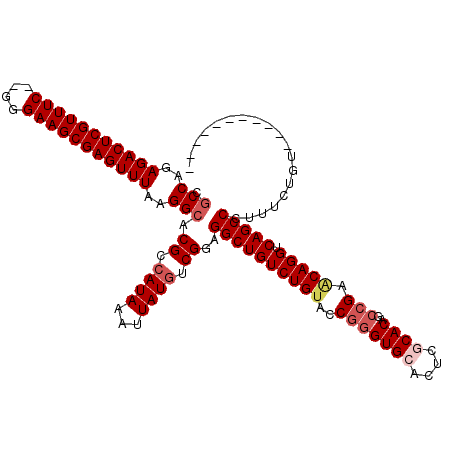
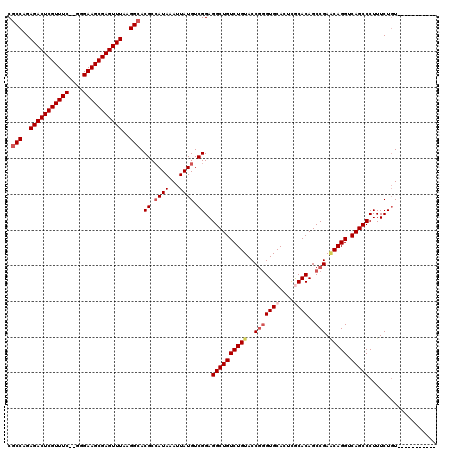
| Location | 1,342,232 – 1,342,332 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.07 |
| Mean single sequence MFE | -34.98 |
| Consensus MFE | -32.12 |
| Energy contribution | -33.12 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.35 |
| SVM RNA-class probability | 0.992828 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 1342232 100 - 20766785 A------------------ACUUCUCAGCACAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCAUAUUAUCCUCCCUAUGCGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACG .------------------.............(((((((....(((.....))))))))))(((((..........)))))(((..((((((((((.--...))))))))))..)))... ( -34.50) >DroSec_CAF1 60577 100 - 1 A------------------ACUUCCCAGCACAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCAUAUUAUCCUCCCUAUGCGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACG .------------------.............(((((((....(((.....))))))))))(((((..........)))))(((..((((((((((.--...))))))))))..)))... ( -34.50) >DroSim_CAF1 38690 100 - 1 A------------------ACUUCCCAGCGCAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCAUAUUAUCCUCCCUAUGCGCCAGAGACUCGUUUC--GGGAAGCGAGUUUAAGGCACG .------------------.............(((((((....(((.....))))))))))(((((..........)))))(((..((((((((((.--...))))))))))..)))... ( -34.50) >DroEre_CAF1 41959 118 - 1 ACUUUCUCAGCUUCUCGCAACUUCUCAGCCCAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCUUAUUAUCCUCCCAAUGCGCCGGAGACUCGUUUC--GGGAAGCGAGUUUAAGGAACG .(((.(((.((((((((.(((.((((.((...(((((((....(((.....))))))))))((................))))..))))...))).)--))))))))))...)))..... ( -37.39) >DroYak_CAF1 29223 102 - 1 A------------------ACUUCUCAGCCCAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCUUAUUAUCCUUCCAAUGCGCCAGAGACUCGUUUCGCGGGAAGCGAGUUUAAGGCACG .------------------.......(((...(((((((....(((.....)))))))))))))...............(.(((..(((((((((((....)))))))))))..))).). ( -34.00) >consensus A__________________ACUUCUCAGCACAGGCCGGAUUUUCGAUUCACUCGUUCGGCCGCAUAUUAUCCUCCCUAUGCGCCAGAGACUCGUUUC__GGGAAGCGAGUUUAAGGCACG ................................(((((((....(((.....))))))))))(((((..........)))))(((..(((((((((((....)))))))))))..)))... (-32.12 = -33.12 + 1.00)

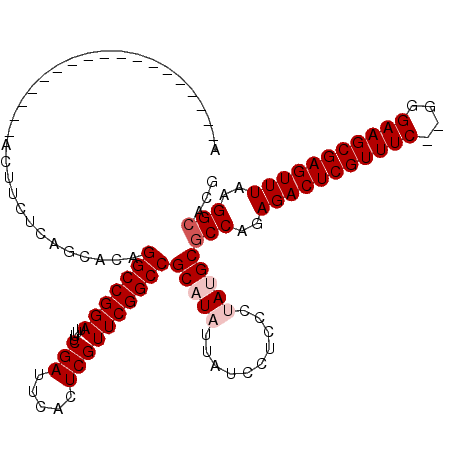
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:26:41 2006