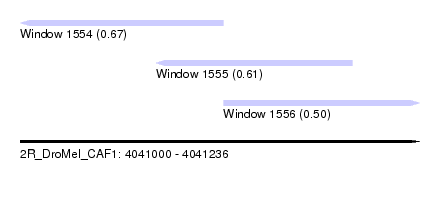
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,041,000 – 4,041,236 |
| Length | 236 |
| Max. P | 0.665915 |

| Location | 4,041,000 – 4,041,120 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.29 |
| Mean single sequence MFE | -28.24 |
| Consensus MFE | -22.76 |
| Energy contribution | -22.20 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
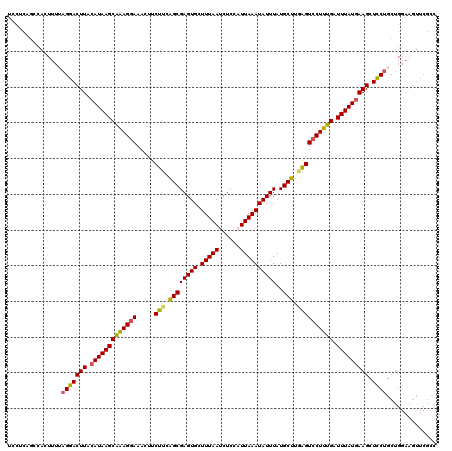
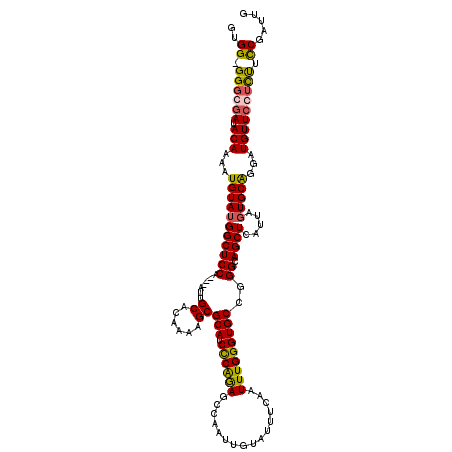
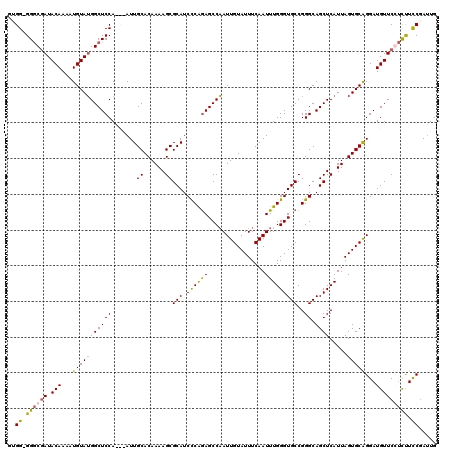
>2R_DroMel_CAF1 4041000 120 - 20766785 UCCUCUGACACUUUUAGGACUUACAUAAGCAGAGGAAACUUCUUCAGCGAGUGCUUUAAUCUCCAUUAAAUAUUUAUGCUUGAGUCCUUUGAUUUAUGAAGCUCCUGCUGGAAGUUCGUU ......((.(((((((((((((.(((((((((((((.....(((.((((((((.((((((....)))))))))))..))).)))))))))).))))))))).)))))...)))))))... ( -29.70) >DroSec_CAF1 23616 119 - 1 UCCUCAGCCAGUUUUAGGACUUACAUAAGCAAAG-AAACUUCUUCGGCGAGUGCUUUAAUCUCCAUUAAAUAUUUAUGCUUGAGUCCUUUGAUUUAUGAAGCUCCUGCUGGAAGUUCGCC .......(((((...(((((((.(((((((((((-......(((.((((((((.((((((....)))))))))))..))).)))..))))).))))))))).)))))))))......... ( -27.40) >DroSim_CAF1 25766 120 - 1 UCCUCAUUCACUUUUAGGACUUACAUAAGCAAAGGAAACUUCUUUAGCGAGUGCUUUAAUCUCCAUUAAAUAUUUAUGCUUGAGUCCUUUGAUUUAUGAAGCUCCUGCUGGAAGUUCGCC .........(((((((((((((.(((((((((((((.....(((.((((((((.((((((....)))))))))))..))).)))))))))).))))))))).)))))...)))))..... ( -28.30) >DroEre_CAF1 23000 105 - 1 ---GCAACCAUCUUGAGUACUUGCAUAAGCAAAGGAAACUUCCGCAGCGAGUGCUUUAAGCUCUAUUAAAUAUUUAUGCUUCGGUCCUUUGAUUUAUGAAGCUGCCGC------------ ---(((.....(((((((((((((....((.(((....)))..)).)))))))).))))).................(((((((((....)))...)))))))))...------------ ( -26.70) >DroYak_CAF1 26652 117 - 1 ---GCAGGCAUUUUGAGGACUUAGAUAAGCGAAGGAAACAUCUUCAGCGAGUGCUUUAAUCUUCCUUAAAUAUUUAUGCUUCAGUCCUUUGAUUUAUGAAGCUCCUGCUGGAUGUUCGUA ---(((((...((((((((...(((((((((..(....).((......)).)))))..)))))))))))).......(((((((((....)))...)))))).)))))............ ( -29.10) >consensus UCCUCAGCCACUUUUAGGACUUACAUAAGCAAAGGAAACUUCUUCAGCGAGUGCUUUAAUCUCCAUUAAAUAUUUAUGCUUGAGUCCUUUGAUUUAUGAAGCUCCUGCUGGAAGUUCGCC ...............(((((((.(((((((((((((.....(((.((((((((.(((((......))))))))))..))).)))))))))).))))))))).)))).............. (-22.76 = -22.20 + -0.56)
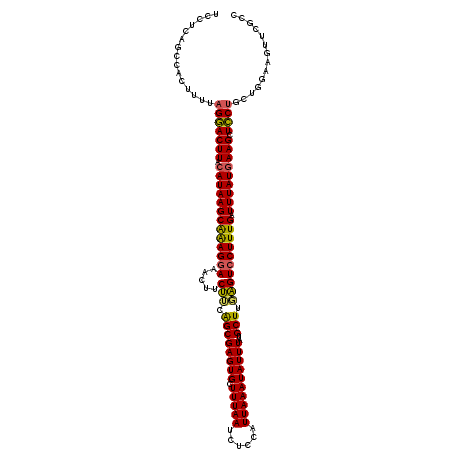


| Location | 4,041,080 – 4,041,196 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.25 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -20.30 |
| Energy contribution | -19.55 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.607272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4041080 116 - 20766785 CACCCAAAUUGAAAUACAAUUGGCUCCGGGAUGCGCUUUUGUGCAAU---UGGAGCCAUACAUUUUGUAUAGCCC-CCACUCCUCUGACACUUUUAGGACUUACAUAAGCAGAGGAAACU .............((((((.(((((((((..(((((....))))).)---))))))))......)))))).....-....(((((((...(((...(......)..)))))))))).... ( -34.50) >DroSec_CAF1 23696 119 - 1 CACCCAAAUUGAAAUACAAUUGGCUCUGGGAUGCGCUUUUGUGCAAUUGUUGGAGCCAUACAUUUUGUAUCGCCCCCCACUCCUCAGCCAGUUUUAGGACUUACAUAAGCAAAG-AAACU ......(((((.....)))))((((..(((((((((....)))))...((.((.((.((((.....)))).))..)).)))))).))))((((((..(.(((....))))...)-))))) ( -27.40) >DroSim_CAF1 25846 119 - 1 CACCCAAAUUGAAAUACAAUUGGCUCUGGGAUGCGCUUUUGUGCAAUUGUUGGAGCCAUACAUUUUGUAUCGCCC-CCUCUCCUCAUUCACUUUUAGGACUUACAUAAGCAAAGGAAACU ...((.....(..((((((.((((((..(.((((((....)))))..).)..))))))......))))))..)..-....((((.(.......).))))..............))..... ( -26.00) >DroYak_CAF1 26732 111 - 1 CACCCAAAUUGAAAUACAAUAGGCUUUGAGAUGCGCUUUUGUGCAAU---UGGAGCCUUACAUUUUGUAUCGGCC-CC-----GCAGGCAUUUUGAGGACUUAGAUAAGCGAAGGAAACA ..((((((.....((((((.((((((..(..(((((....))))).)---..))))))......))))))..(((-..-----...)))..)))).))...............(....). ( -30.70) >consensus CACCCAAAUUGAAAUACAAUUGGCUCUGGGAUGCGCUUUUGUGCAAU___UGGAGCCAUACAUUUUGUAUCGCCC_CCACUCCUCAGACACUUUUAGGACUUACAUAAGCAAAGGAAACU .............((((((.((((((((...(((((....))))).....))))))))......)))))).................................................. (-20.30 = -19.55 + -0.75)

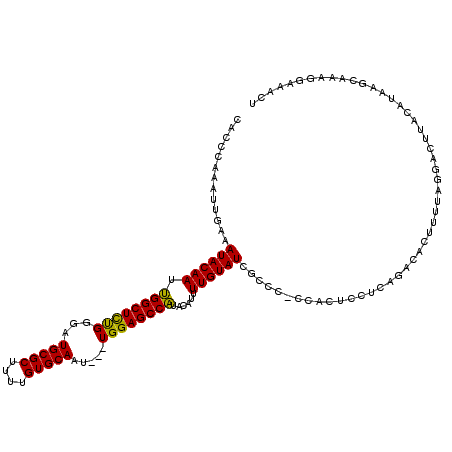
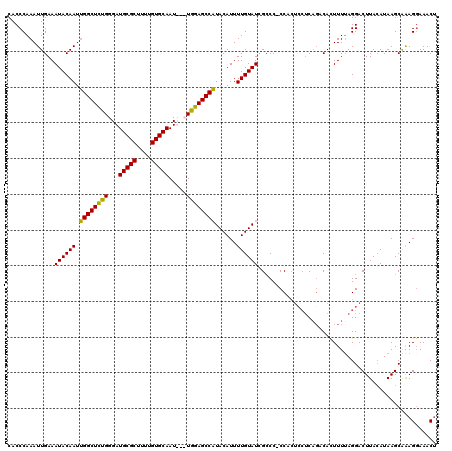
| Location | 4,041,120 – 4,041,236 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.34 |
| Mean single sequence MFE | -37.52 |
| Consensus MFE | -27.90 |
| Energy contribution | -27.84 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4041120 116 + 20766785 GUGG-GGGCUAUACAAAAUGUAUGGCUCCA---AUUGCACAAAAGCGCAUCCCGGAGCCAAUUGUAUUUCAAUUUGGGUGCCGGGCAGCUCAUUAGUGCAGGAUGUUCCUCCUUCGAUUG ...(-(((((((((.....)))))))))).---..(((((.((.(.((..(((((.(((.((((.....))))...))).)))))..)).).)).)))))(((......)))........ ( -43.20) >DroSec_CAF1 23735 120 + 1 GUGGGGGGCGAUACAAAAUGUAUGGCUCCAACAAUUGCACAAAAGCGCAUCCCAGAGCCAAUUGUAUUUCAAUUUGGGUGCCGGGCAGCUCAUUAGUGCAGGAUGUUCGUCUUCCGAUUG .((((((((.((((.....)))).)))))((((.((((((.((.(.((..(((.(.(((.((((.....))))...))).).)))..)).).)).))))))..))))......))).... ( -37.70) >DroSim_CAF1 25886 119 + 1 GAGG-GGGCGAUACAAAAUGUAUGGCUCCAACAAUUGCACAAAAGCGCAUCCCAGAGCCAAUUGUAUUUCAAUUUGGGUGCCGGGCAGCUCAUUAGUGCAGGAUGUUCGUCUUCCGAUUG ...(-((((.((((.....)))).)))))..(((((((((.((.(.((..(((.(.(((.((((.....))))...))).).)))..)).).)).)))).(((.(.....).)))))))) ( -36.30) >DroYak_CAF1 26769 114 + 1 --GG-GGCCGAUACAAAAUGUAAGGCUCCA---AUUGCACAAAAGCGCAUCUCAAAGCCUAUUGUAUUUCAAUUUGGGUGCCGGGCAGCUCAUUAGUGCGGGAUGUUCCAUUUCCUAAUG --((-((((..(((.....))).)))))).---...((......))(((((((...((((...((((((......)))))).)))).((........))))))))).............. ( -32.90) >consensus GUGG_GGGCGAUACAAAAUGUAUGGCUCCA___AUUGCACAAAAGCGCAUCCCAGAGCCAAUUGUAUUUCAAUUUGGGUGCCGGGCAGCUCAUUAGUGCAGGAUGUUCCUCUUCCGAUUG ..((.((((((.(((...(((((((((((.......((......))(((.((((((................)))))))))..)).)))).....)))))...))))))))).))..... (-27.90 = -27.84 + -0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:50:15 2006