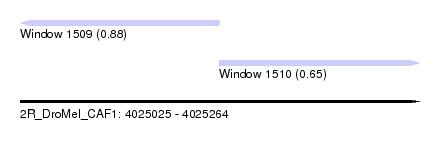
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,025,025 – 4,025,264 |
| Length | 239 |
| Max. P | 0.876938 |

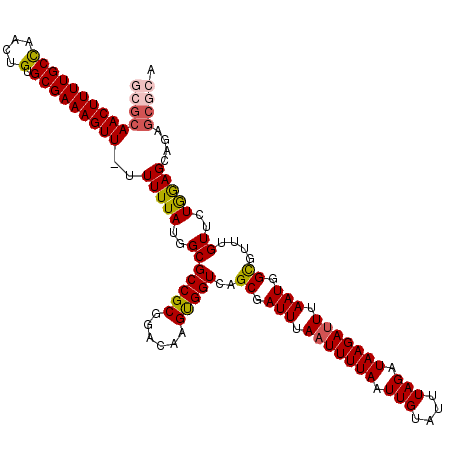
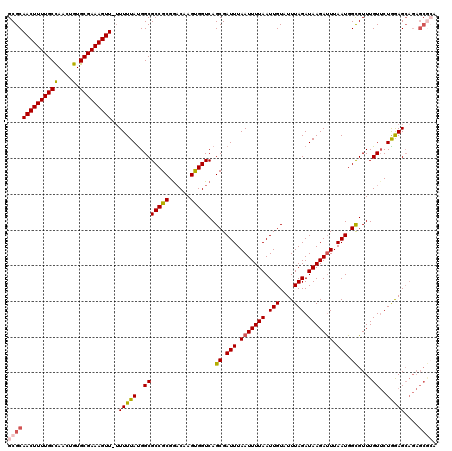
| Location | 4,025,025 – 4,025,144 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.13 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -27.26 |
| Energy contribution | -27.70 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.876938 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4025025 119 - 20766785 GCGCAACUUUUGCCAACUGUGCGAAAGUU-UUUUUAUGGCGCCGCGAACAAGCGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGAUUUAAUGGCGUUUGUUCUAGAGCAGAGCGCA ((((((((((((((....).)))))))))-.(((((..(((((((......)))))..((.(((.(((((((.(((....))).))))))).))).))....))..)))))....)))). ( -36.10) >DroSec_CAF1 7828 119 - 1 GCGCAACUUUUGCCAACUGUGCGAAAGUU-UUUUUAUGGCGCCGCGGACAAGUGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGAUUUAAUGGCGUUUGUUCUGGAGCAGAGCGCA ((((((((((((((....).)))))))))-........((.(((.(((((((......((.(((.(((((((.(((....))).))))))).))).)).)))))))))).))...)))). ( -35.50) >DroSim_CAF1 9878 119 - 1 GCGCAACUUUUGCCAACUGUGCGAAAGUU-UUUUUAUGGCGCCGCGGACAAGUGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGAUUUAAUGGCGUUUGUUCUGGAGCAGAGCGCA ((((((((((((((....).)))))))))-........((.(((.(((((((......((.(((.(((((((.(((....))).))))))).))).)).)))))))))).))...)))). ( -35.50) >DroEre_CAF1 7080 111 - 1 GCGCAACUUUUGCCAACUGUGCGAAAGUUGUUUUUACGGCGCCGCGGCCAAGUGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGACUUAAUGGUGUUUGUGCUGGAG--------- ..((((((((((((....).))))))))))).....(((((((((((((....)))).))).........................((((........))))))))))...--------- ( -32.40) >DroYak_CAF1 9625 118 - 1 GCGCAACUUUUGCUAACUGUGCGAAAGUU--UUUUAUGGCGCCGCGGACAAGUGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGAUUUAAUGGUGUUUGUUCUAAAGCAGAGCAAA (((((((((((((.......)))))))))--.......))))(((.(((.....))).)))(((.(((((((.(((....))).))))))).)))....((((((((.....)))))))) ( -29.51) >consensus GCGCAACUUUUGCCAACUGUGCGAAAGUU_UUUUUAUGGCGCCGCGGACAAGUGGUCAGCGAUUUAAUUUUAAUUGUAUUUAGAUAAGAUUUAAUGGCGUUUGUUCUGGAGCAGAGCGCA ((((((((((((((....).)))))))))..(((((..(((((((......)))))..((.(((.(((((((.(((....))).))))))).))).))....))..)))))....)))). (-27.26 = -27.70 + 0.44)

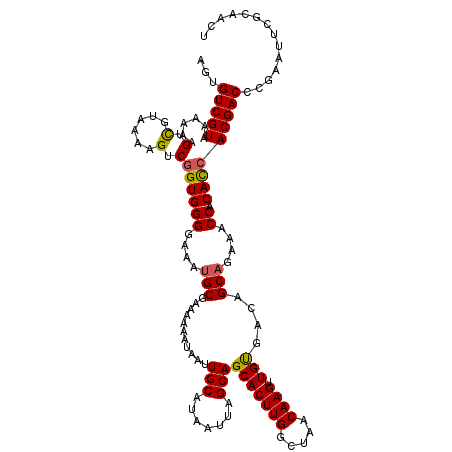
| Location | 4,025,144 – 4,025,264 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.17 |
| Mean single sequence MFE | -27.19 |
| Consensus MFE | -24.72 |
| Energy contribution | -24.64 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.653823 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4025144 120 + 20766785 AGUGUCGUAAAAAAGUCGUAAAAGUCGGUGGGGAAAUGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGUGACAGCAGAAACCACACCACGACCCGAAUUCGCAACU .(.(((((......(.(......).)((((((....(((...........(((.......)))(((((((.....)))).)))....)))....)).))))))))).)............ ( -27.30) >DroSec_CAF1 7947 120 + 1 AGUGUCGUAAAAAAGUCGUAAAAGUCGGUGGGGAAAGGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGUGACAGCAGAAACCACACCACGACCCGAAUCCGCAACU .(.(((((......(.(......).)((((((.....((...........(((.......)))(((((((.....)))).)))....)).....)).))))))))).)............ ( -25.70) >DroSim_CAF1 9997 120 + 1 AGUGUCGUAAAAAAGUCGUAAAAGUCGGUGGGGAAAUGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGUGACAGCAGAAACCACACCACGACCCGAAUUCGCAACU .(.(((((......(.(......).)((((((....(((...........(((.......)))(((((((.....)))).)))....)))....)).))))))))).)............ ( -27.30) >DroEre_CAF1 7191 120 + 1 AAUGUCGUAAAAAAGUUGUAAAAGUCGGUGGGGAAAUGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGCGACAGCAGAAACCACACCACGACCCCAAUUCGCAGCU .............((((((....((.((((((....(((...........(((.......)))(((((((.....)))).)))....)))....)).))))))((.......)))))))) ( -28.30) >DroYak_CAF1 9743 120 + 1 AGUGUCGUAAAAAAGUCGUAAAAGUCUGUGGGGAAAUGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGUGACAGCAGUAACCACAUCACGACCCCAAUUCGCCACU ((((.((.......(((((.......(((((....((((((........)))))).....((.(((((((.....)))).)))....)).....)))))..))))).......)).)))) ( -27.34) >consensus AGUGUCGUAAAAAAGUCGUAAAAGUCGGUGGGGAAAUGCGAAAAAAUAAUUGCAUAAUUAGCAGCACUUGGCUAACAAGUUGUGACAGCAGAAACCACACCACGACCCGAAUUCGCAACU ...(((((......(.(......).)((((((....(((...........(((.......)))(((((((.....)))).)))....)))....)).))))))))).............. (-24.72 = -24.64 + -0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:29 2006