
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 3,875,180 – 3,875,298 |
| Length | 118 |
| Max. P | 0.999583 |

| Location | 3,875,180 – 3,875,298 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 80.43 |
| Mean single sequence MFE | -25.90 |
| Consensus MFE | -18.54 |
| Energy contribution | -19.60 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.866859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 3875180 118 + 20766785 UUUUUUGUUAGGAUCGGAUACCUGACUGCUAAAGCAAAUUAAUAGAAACAUCACUUUGUAUCUAUUACUUCGUUUUAAAAGCCAGACCCUCAACUUGCACUUUUACGUGUUUCGGUAG ............((((((...(((.((..((((((.((.(((((((.(((......))).))))))).)).))))))..)).)))...........((((......)))))))))).. ( -24.40) >DroSec_CAF1 2152 117 + 1 UUU-UUAUUAAGAUAGGAGACCUGACCGCUAAAGCAAAUUAAUAGAAACAUAUAUUUGUAUCUAUUACUUUGUUUUAAAAGCCAGACCCUCAACUUGCACUUCUACGUGUAUCGUUAG ...-............(((..(((.(...(((((((((.(((((((.(((......))).))))))).)))))))))...).)))...)))(((.(((((......)))))..))).. ( -25.70) >DroEre_CAF1 2365 116 + 1 UUG-UUGUACGGAUCGAAGACCUGACUUCUAAAGCAAAGUAAUAGAAACGC-AACUUGUAUCUAUUGCUUCGUUUUAAAAGGCAGACCCUCCGCUUCCAUUCUUUCAGAUUUAGUUAA ...-......(((.((.((..(((.(((.((((((.((((((((((.(((.-....))).)))))))))).)))))).))).)))...)).))..))).................... ( -27.70) >DroYak_CAF1 2424 117 + 1 UUG-UUGCUAGGAUCGAAGACCUGACUUCUAAAGCAAAGUAAUAGAAACGCCAACUUGCAUCUAUUACUUCGUUUUAAAAGGCAGUCCCUCAACUUCCAUUUUUUCAGAUUUGUGUAU ...-......(((..((.(..(((.(((.((((((.((((((((((...((......)).)))))))))).)))))).))).)))..).))....))).................... ( -25.80) >consensus UUG_UUGUUAGGAUCGAAGACCUGACUGCUAAAGCAAAGUAAUAGAAACACCAACUUGUAUCUAUUACUUCGUUUUAAAAGCCAGACCCUCAACUUCCACUUUUACAGAUUUCGGUAG .....................(((.(((.((((((.((((((((((.(((......))).)))))))))).)))))).))).)))................................. (-18.54 = -19.60 + 1.06)

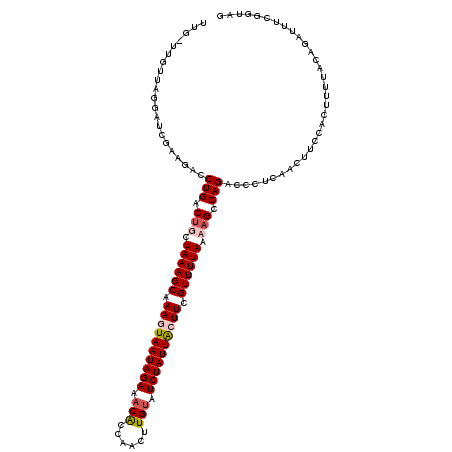
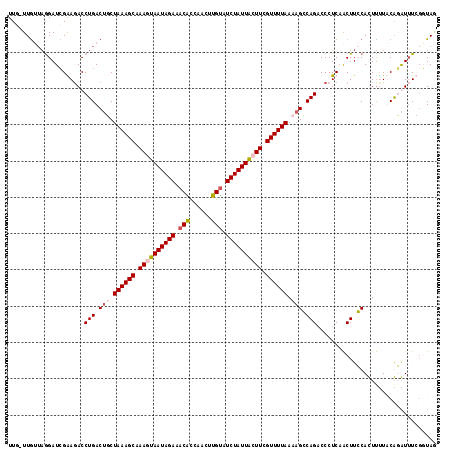
| Location | 3,875,180 – 3,875,298 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 80.43 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -27.11 |
| Energy contribution | -29.05 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.43 |
| Structure conservation index | 0.82 |
| SVM decision value | 3.75 |
| SVM RNA-class probability | 0.999583 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 3875180 118 - 20766785 CUACCGAAACACGUAAAAGUGCAAGUUGAGGGUCUGGCUUUUAAAACGAAGUAAUAGAUACAAAGUGAUGUUUCUAUUAAUUUGCUUUAGCAGUCAGGUAUCCGAUCCUAACAAAAAA .........(((......)))...(((.(((((((((((.(((((.((((.(((((((.(((......))).))))))).)))).))))).)))))((...)))))))))))...... ( -29.80) >DroSec_CAF1 2152 117 - 1 CUAACGAUACACGUAGAAGUGCAAGUUGAGGGUCUGGCUUUUAAAACAAAGUAAUAGAUACAAAUAUAUGUUUCUAUUAAUUUGCUUUAGCGGUCAGGUCUCCUAUCUUAAUAA-AAA .....(((((((......)))......(..(..((((((.(((((.((((.(((((((.(((......))).))))))).)))).))))).))))))..)..))))).......-... ( -30.90) >DroEre_CAF1 2365 116 - 1 UUAACUAAAUCUGAAAGAAUGGAAGCGGAGGGUCUGCCUUUUAAAACGAAGCAAUAGAUACAAGUU-GCGUUUCUAUUACUUUGCUUUAGAAGUCAGGUCUUCGAUCCGUACAA-CAA ................(.(((((....((((..(((.((((((((.(((((.((((((.((.....-..)).)))))).))))).)))))))).)))..))))..))))).)..-... ( -34.20) >DroYak_CAF1 2424 117 - 1 AUACACAAAUCUGAAAAAAUGGAAGUUGAGGGACUGCCUUUUAAAACGAAGUAAUAGAUGCAAGUUGGCGUUUCUAUUACUUUGCUUUAGAAGUCAGGUCUUCGAUCCUAGCAA-CAA ....................(((..((((((..(((.((((((((.(((((((((((((((......)))..)))))))))))).)))))))).)))..)))))))))......-... ( -37.10) >consensus CUAACGAAACACGAAAAAAUGCAAGUUGAGGGUCUGCCUUUUAAAACGAAGUAAUAGAUACAAAUUGACGUUUCUAUUAAUUUGCUUUAGAAGUCAGGUCUCCGAUCCUAACAA_AAA ...........................((((..((((((((((((.((((((((((((.((........)).)))))))))))).))))))))))))..))))............... (-27.11 = -29.05 + 1.94)

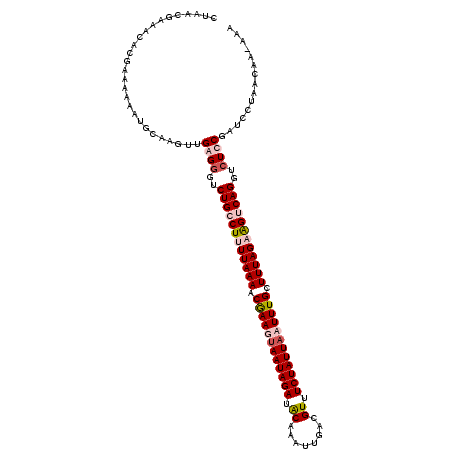
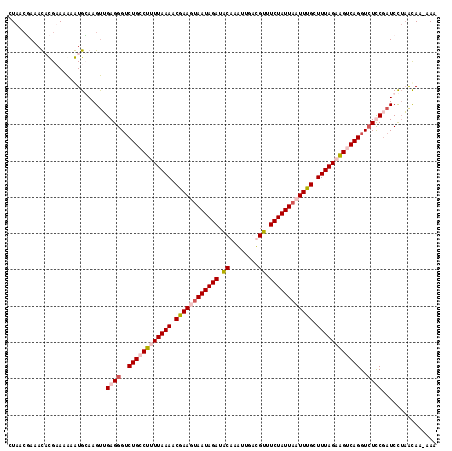
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:48:20 2006