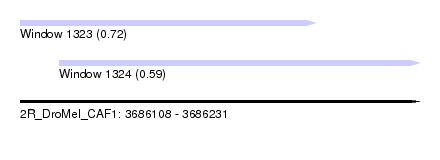

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 3,686,108 – 3,686,231 |
| Length | 123 |
| Max. P | 0.718951 |

| Location | 3,686,108 – 3,686,199 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 94.84 |
| Mean single sequence MFE | -25.89 |
| Consensus MFE | -20.96 |
| Energy contribution | -20.68 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.718951 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 3686108 91 + 20766785 UGUGUUUGGCGGAUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCAGCGAAUUCUGAGCGCGUAAA .((((((............(((((((((((.....)))))...))))))((((.((.(((((....))))).)).)))).))))))..... ( -26.80) >DroSec_CAF1 6392 91 + 1 UGUGUUUGGCGGAUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCUGCGAAUUCUGAGCGCGCAAA ....(((((((....((((..((((((((((..((((((((.............)))))).))..)))))))))).))))...))).)))) ( -27.02) >DroSim_CAF1 5883 91 + 1 UGUGUUUGGCGGAUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCUGCGAAUUCUGAGCGCGCAAA ....(((((((....((((..((((((((((..((((((((.............)))))).))..)))))))))).))))...))).)))) ( -27.02) >DroEre_CAF1 6454 91 + 1 UGUGUUUGGCGGAUCAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGCUUGAGUGCGGUGUACUAAUAUGCAGCAAAUUCUGAGCGCGCAAA ....(((((((..((((((....((.(((((..((.(((....(..(....)..)...)))))..))))).))...)))))).))).)))) ( -26.80) >DroYak_CAF1 6334 90 + 1 UGUGAUUGGCGGAU-GGAAAAUUGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUAUGCAGCGAAUGCUGAGCGCGCAAA ((((.(..(((..(-....).((((.(((((..((((((((.............)))))).))..))))).)))).)))..)...)))).. ( -21.82) >consensus UGUGUUUGGCGGAUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCAGCGAAUUCUGAGCGCGCAAA ........((((...((((..((((.(((((..((((((((.............)))))).))..))))).)))).))))...).)))... (-20.96 = -20.68 + -0.28)
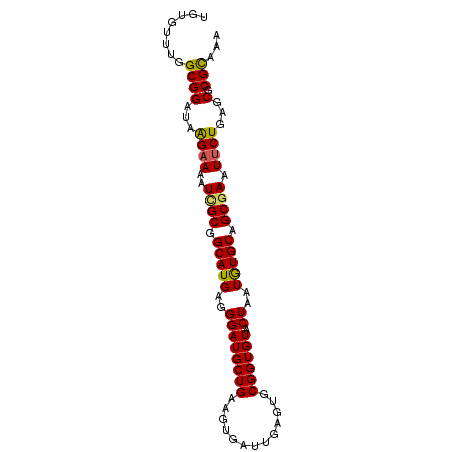


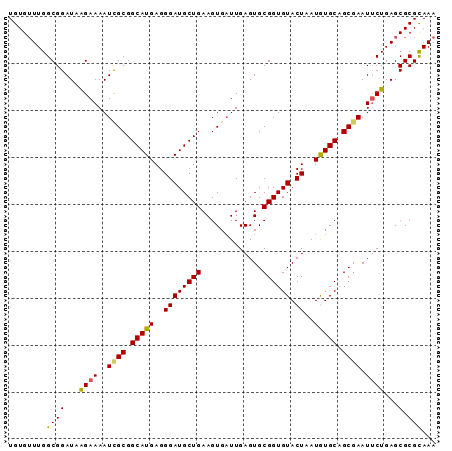
| Location | 3,686,120 – 3,686,231 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 94.32 |
| Mean single sequence MFE | -27.50 |
| Consensus MFE | -21.52 |
| Energy contribution | -21.80 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 3686120 111 + 20766785 AUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCAGCGAAUUCUGAGCGCGUAAAGCGUUUCGAAAAAUUAAACAAACAAAAUGCAU .......(((((((((((.....)))))...))))))..(((((.(((((....))))).)...(((.((((((.....)))))).)))..................)))) ( -29.50) >DroSec_CAF1 6404 111 + 1 AUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCUGCGAAUUCUGAGCGCGCAAAGCGUUUCGAAAAAGUAAACAAACAAAAUGCAU .........((((((((((..((((((((.............)))))).))..)))))))))).(((.((((((.....)))))).)))...................... ( -29.12) >DroSim_CAF1 5895 111 + 1 AUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCUGCGAAUUCUGAGCGCGCAAAACGUUUCGAAAAAGUAAACAAACAAAAUGCAU .......(((((((((((.....)))))...))))))..(((((.(((.....(((((.((.........)))))))....(((((......).))))..)))...))))) ( -24.50) >DroEre_CAF1 6466 111 + 1 AUCAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGCUUGAGUGCGGUGUACUAAUAUGCAGCAAAUUCUGAGCGCGCAAAGCGUUUCAAGAAAGUAAACAAACAAAAUGCAU ..............((((..(..(((((...((((((((((..(.((((......)))).)..)))).))))))....)))))..)......((......))...)))).. ( -28.60) >DroYak_CAF1 6346 110 + 1 AU-GGAAAAUUGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUAUGCAGCGAAUGCUGAGCGCGCAAAGCGUUUCGAAAAAGUAAACAAACAAAAUGCUU ..-..........(((((..(..(((((...(((.((.(((.((.((((......)))).))...))).)))))....)))))..)......((......))...))))). ( -25.80) >consensus AUAAGAAAAUCGCGGCAUGAGGGAUGCUGAAGUGAUUGAGUGCGGUGUACUAAUGUGCAGCGAAUUCUGAGCGCGCAAAGCGUUUCGAAAAAGUAAACAAACAAAAUGCAU .........((((.(((((..((((((((.............)))))).))..))))).)))).(((.((((((.....)))))).)))...................... (-21.52 = -21.80 + 0.28)

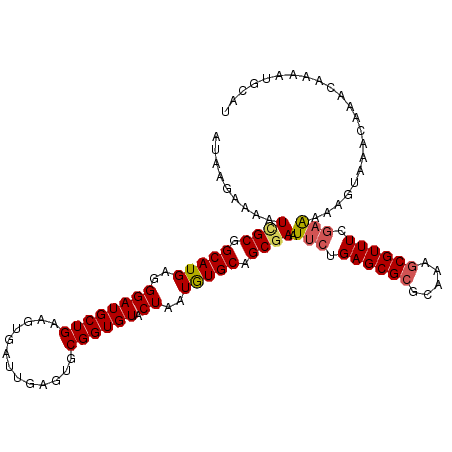
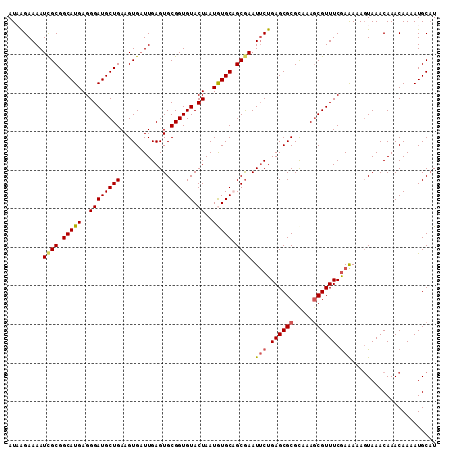
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:29 2006