
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 20,538,555 – 20,538,661 |
| Length | 106 |
| Max. P | 0.699940 |

| Location | 20,538,555 – 20,538,661 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 79.79 |
| Mean single sequence MFE | -30.38 |
| Consensus MFE | -23.39 |
| Energy contribution | -23.12 |
| Covariance contribution | -0.27 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.699940 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
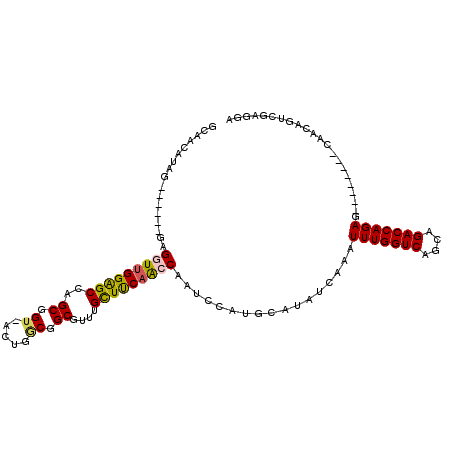
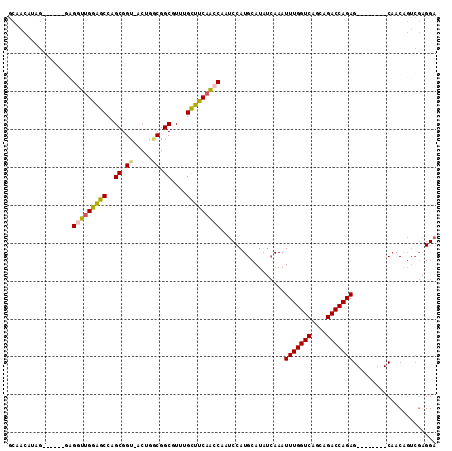
>2R_DroMel_CAF1 20538555 106 - 20766785 GCAACAUAG------GAGGUUGGAGCCAGCGGU-ACUGUCGGCUUUUGCUUCAACCAAUCCAUGCAUAUCAAAUUUGGUCAGCAGACCAGAGCAACAGAGCAACAGUCGAGGA ((..(((.(------(((((((((((.(((.(.-.....).)))...)))))))))..)))))).........(((((((....))))))))).................... ( -32.00) >DroSec_CAF1 7342 98 - 1 GCAACAUAG------GAGGUUGGGGCCAGCGGU-ACUGGCGGCGUUUGCUCCAACCAAUCCAUGCAUAUCAAAUUUGGUCAAUAGACCAGAG--------CAACAGUCGAGGA ((..(((.(------(((((((((((..((.((-....)).))....)))))))))..)))))).........(((((((....))))))))--------)............ ( -34.80) >DroSim_CAF1 3173 98 - 1 GCAACAUAG------GAGGUUGGGGCCAGCGGU-ACUGGCGGCGUUUGCUCCAACCAAUCCAUGCAUAUCAAAUUUGGUCAACAGACCAGAG--------CAACAGUCGAGGA ((..(((.(------(((((((((((..((.((-....)).))....)))))))))..)))))).........(((((((....))))))))--------)............ ( -34.80) >DroEre_CAF1 7182 87 - 1 -----------------GGUUGGAGCCUGCGGU-ACUGACGGCGUUUGUUUCCACCAAUCCAUGCAUAUCAAAUUUGGUCAGCAGACCAGAG--------CAACUGUCGAGGA -----------------(((.(((((.(((.((-....)).)))...))))).)))..(((.((.(((.....(((((((....))))))).--------....))))).))) ( -23.80) >DroYak_CAF1 3239 104 - 1 GCAACAGAGGUUGGAGAGGUUGGAGCCUGCGGU-ACUGGCGGCGUUUGUUUCAAGCAAUCCAUGCAUAUCAAAUUUGGUCAGCAGACCAGAG--------CAACAGUCGAGGA ((....((.((((((..(.(((((((.(((.((-....)).)))...))))))).)..)))).))...))...(((((((....))))))))--------)............ ( -30.40) >DroPer_CAF1 4018 85 - 1 -------------AAAGGGUUGGAGCCUGCGGUCACGGGCGGCUUUUGUUUCAGGCAAGCCAUGCAUA--AAAUUUGGUCAGCAGACCAGAG--------CCACA-----GGA -------------............((((((....))(((((((..(((.....))))))).......--...(((((((....))))))))--------)).))-----)). ( -26.50) >consensus GCAACAUAG______GAGGUUGGAGCCAGCGGU_ACUGGCGGCGUUUGCUUCAACCAAUCCAUGCAUAUCAAAUUUGGUCAGCAGACCAGAG________CAACAGUCGAGGA .................(((((((((..((.((.....)).))....))))))))).................(((((((....)))))))...................... (-23.39 = -23.12 + -0.27)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:14:06 2006