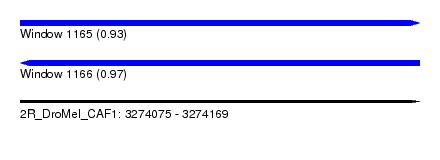
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 3,274,075 – 3,274,169 |
| Length | 94 |
| Max. P | 0.973629 |

| Location | 3,274,075 – 3,274,169 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 95.21 |
| Mean single sequence MFE | -18.88 |
| Consensus MFE | -15.26 |
| Energy contribution | -15.82 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928870 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
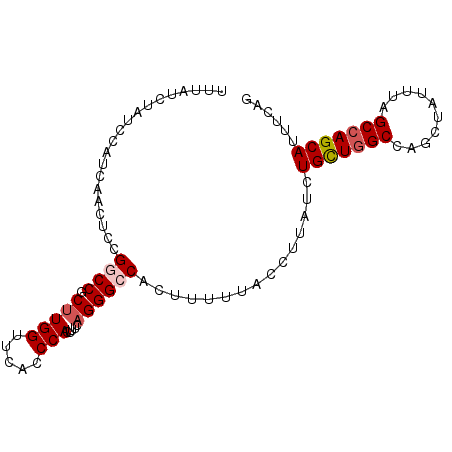
>2R_DroMel_CAF1 3274075 94 + 20766785 UUUAUCUAUCCAUCAACUCCGGCCGCUUGGUUCACCCACUUUCAGGGCCACUUUUUACCUUAUCUGCUGGCCAGCUAUUUAGCCAGCAUUUCAG ....................((((.(((((.....))).....))))))...............(((((((..........)))))))...... ( -20.50) >DroSec_CAF1 42707 94 + 1 UUUAUCUAUCCAUCAACUCCGUCCGCUUGGUUCACCCACUUUCAGGGCCACUUUUUACCUUAUCUGUUGGCCAGCUAUUUAGCCAGCAUUUCAG ....................((((....(((......)))....))))................(((((((..........)))))))...... ( -14.90) >DroSim_CAF1 53908 94 + 1 UUUAUCUAUCCAUCAACUCCGUCCGCUUGGUUCACCCACUUUCAGGGCCACUUUUUACCUUAUCUGUUGGCCAGCUAUUUAGCCAGCAUUUCAG ....................((((....(((......)))....))))................(((((((..........)))))))...... ( -14.90) >DroEre_CAF1 41859 94 + 1 UUUAUCUAUCCAUCAACUCUGGCCGCUUGGUUCACCCACUUUCAGGGCCACUUUUUACCUUAUCUGCUGGCCAGCUAUAUGGCCAGCAUUUCAG ...................(((((.(((((.....))).....)))))))..............(((((((((......)))))))))...... ( -27.30) >DroYak_CAF1 41880 94 + 1 UUUAUCUAUCCAUCAACUCUGGCCGCAUGGUUAACCCACUUUCAGGGCCACUUUUUACCUUAUCUGCUUGCUAGCUAUUUUGCCAGCAUUUCAG ...................(((((....(((......))).....))))).............(((..((((.((......)).))))...))) ( -16.80) >consensus UUUAUCUAUCCAUCAACUCCGGCCGCUUGGUUCACCCACUUUCAGGGCCACUUUUUACCUUAUCUGCUGGCCAGCUAUUUAGCCAGCAUUUCAG ....................((((.(((((.....))).....))))))...............(((((((..........)))))))...... (-15.26 = -15.82 + 0.56)

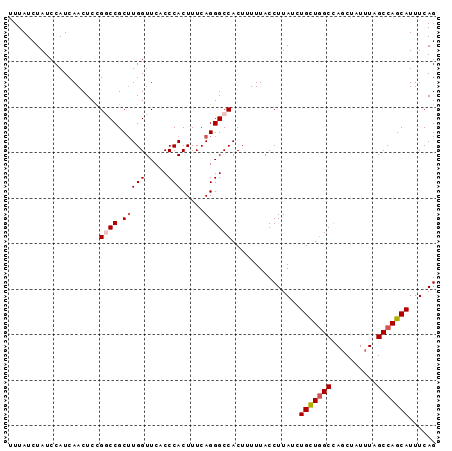
| Location | 3,274,075 – 3,274,169 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 95.21 |
| Mean single sequence MFE | -25.06 |
| Consensus MFE | -20.30 |
| Energy contribution | -21.18 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.973629 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
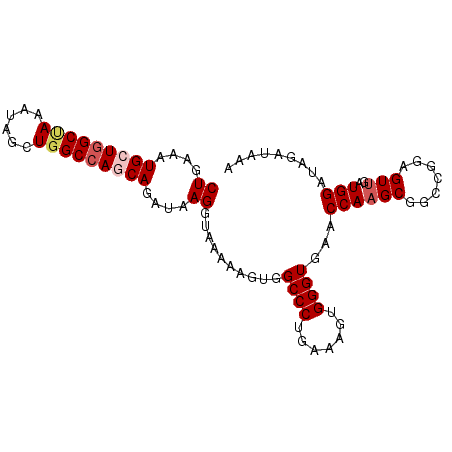
>2R_DroMel_CAF1 3274075 94 - 20766785 CUGAAAUGCUGGCUAAAUAGCUGGCCAGCAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUGAACCAAGCGGCCGGAGUUGAUGGAUAGAUAAA ((....(((((((((......)))))))))....))........(((((((.......((....)).)).)))))................... ( -27.30) >DroSec_CAF1 42707 94 - 1 CUGAAAUGCUGGCUAAAUAGCUGGCCAACAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUGAACCAAGCGGACGGAGUUGAUGGAUAGAUAAA (((....((..(((....)))..))((((......(((........)))(((....(((.....)))..))).....)))).....)))..... ( -20.30) >DroSim_CAF1 53908 94 - 1 CUGAAAUGCUGGCUAAAUAGCUGGCCAACAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUGAACCAAGCGGACGGAGUUGAUGGAUAGAUAAA (((....((..(((....)))..))((((......(((........)))(((....(((.....)))..))).....)))).....)))..... ( -20.30) >DroEre_CAF1 41859 94 - 1 CUGAAAUGCUGGCCAUAUAGCUGGCCAGCAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUGAACCAAGCGGCCAGAGUUGAUGGAUAGAUAAA ((....(((((((((......)))))))))....))........(((((((.......((....)).)).)))))................... ( -30.10) >DroYak_CAF1 41880 94 - 1 CUGAAAUGCUGGCAAAAUAGCUAGCAAGCAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUUAACCAUGCGGCCAGAGUUGAUGGAUAGAUAAA (((...(((((((......)))))))..))).............((((((.......)))))).((((.((((....))))))))......... ( -27.30) >consensus CUGAAAUGCUGGCUAAAUAGCUGGCCAGCAGAUAAGGUAAAAAGUGGCCCUGAAAGUGGGUGAACCAAGCGGCCGGAGUUGAUGGAUAGAUAAA ((....(((((((((......)))))))))....))..........((((.......))))...((((((.......)))..)))......... (-20.30 = -21.18 + 0.88)

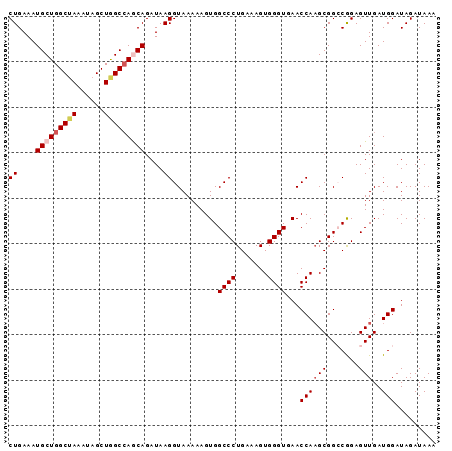
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:43:53 2006