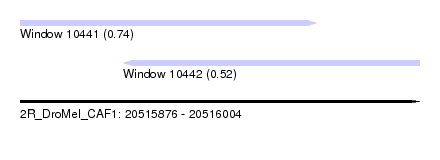
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 20,515,876 – 20,516,004 |
| Length | 128 |
| Max. P | 0.739006 |

| Location | 20,515,876 – 20,515,971 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 86.82 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -15.09 |
| Energy contribution | -15.15 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.739006 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
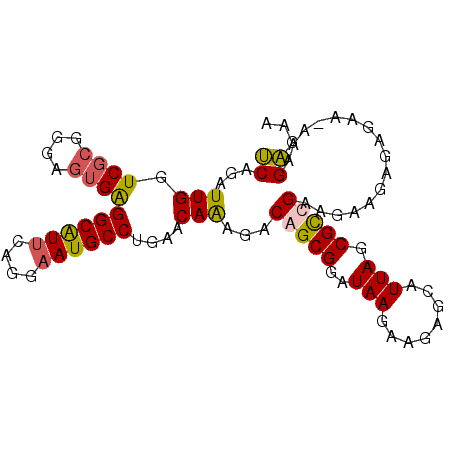
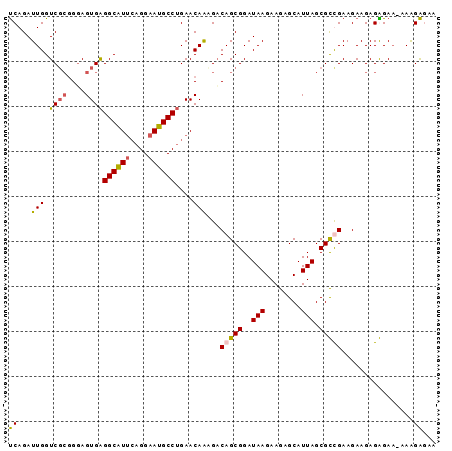
>2R_DroMel_CAF1 20515876 95 + 20766785 UCAGAUUGGUCUCGGGAGUGAGGCGUUCAGGAAUGCCUGAACAAAGACAGCGGAUAAGAAGAGCAUUAGCGCUGAAGAAGAGGGAAAAAAGAAAA ........((((((....))))))(((((((....))))))).....(((((..(((........))).)))))..................... ( -24.70) >DroSec_CAF1 17485 94 + 1 UCAGAUUGGUCGCGGUAGUGAGGCAUUCAGGAAUGCCUGAACAAAGACAGCGGAUAAGAAGAGCAUUAGCGUCGAAGAAGAGAGAA-AAAGAGAU ((...(((..(((.((.((.(((((((....)))))))..))....)).)))..)))((...((....)).))))...........-........ ( -18.20) >DroSim_CAF1 17453 94 + 1 UCAGAUUGGUCGCGGGAGUGAGGCAUUCAGGAAUGCCUGAACAAAGACAGCGGAUAAGAAGAGCAUUAGCGUCGAAGAAGAGAGAA-AAAGAGAA ((...((.(.(((....((.(((((((....)))))))..)).......((...........))....))).).))...)).....-........ ( -18.00) >DroEre_CAF1 17563 95 + 1 CCAGAUUGGUCGGCGGAGUGGGGCAUGGAGGAAUGCCAGAACAGCGUCAGCGGAUAAGAAGAGCAUUAGCGCUGAAGAAGAGGGACAUAGGGGAG ((......(((.((...((..(((((......)))))...)).)).((((((..(((........))).))))))........))).....)).. ( -25.00) >consensus UCAGAUUGGUCGCGGGAGUGAGGCAUUCAGGAAUGCCUGAACAAAGACAGCGGAUAAGAAGAGCAUUAGCGCCGAAGAAGAGAGAA_AAAGAGAA ((...(((.((((....))))((((((....))))))....)))...(((((..(((........))).)))))................))... (-15.09 = -15.15 + 0.06)

| Location | 20,515,909 – 20,516,004 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 84.82 |
| Mean single sequence MFE | -18.90 |
| Consensus MFE | -13.45 |
| Energy contribution | -13.32 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.71 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
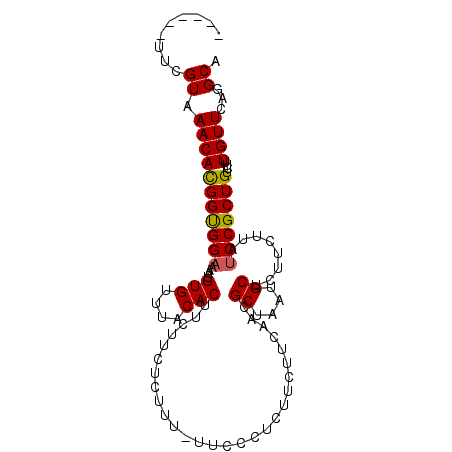
>2R_DroMel_CAF1 20515909 95 - 20766785 ------UUCGUGAACACGGUGGA-AAUGUGUUUACACCUCUUUUCUUUUUUCCCUCUUCUUCAGCGCUAAUGCUCUUCUUAUCCGCUGUCUUUGUUCAGGCA ------..(.(((((((((((((-...(((....))).........................((.((....)).)).....))))))))....))))).).. ( -19.50) >DroSec_CAF1 17518 95 - 1 ------UUCGUAAACACGGCGGAAAAUGUGUUUACACUUCAUCUCUUU-UUCUCUCUUCUUCGACGCUAAUGCUCUUCUUAUCCGCUGUCUUUGUUCAGGCA ------....((((.((((((((....(((....)))...........-.............((.((....))))......)))))))).))))........ ( -18.40) >DroSim_CAF1 17486 95 - 1 ------UUCGUAAACACGGUGGAAAAUGUGUUUACACUUCUUCUCUUU-UUCUCUCUUCUUCGACGCUAAUGCUCUUCUUAUCCGCUGUCUUUGUUCAGGCA ------....((((.((((((((....(((....)))...........-.............((.((....))))......)))))))).))))........ ( -16.50) >DroEre_CAF1 17596 101 - 1 UGAGUAUUCGUAAACAUGGUGGCAA-AGUGUUUACACUUCCUCCCCUAUGUCCCUCUUCUUCAGCGCUAAUGCUCUUCUUAUCCGCUGACGCUGUUCUGGCA .............((((((.((..(-((((....)))))...)).)))))).........((((((.(((........)))..)))))).(((.....))). ( -21.20) >consensus ______UUCGUAAACACGGUGGAAAAUGUGUUUACACUUCUUCUCUUU_UUCCCUCUUCUUCAACGCUAAUGCUCUUCUUAUCCGCUGUCUUUGUUCAGGCA .........((.(((((((((((....(((....)))............................((....))........)))))))....))))...)). (-13.45 = -13.32 + -0.12)

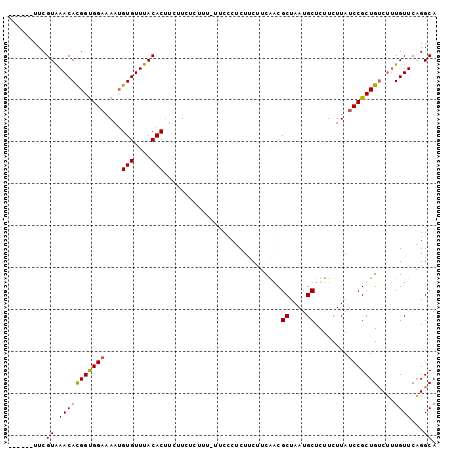
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:13:48 2006