
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 3,199,877 – 3,199,968 |
| Length | 91 |
| Max. P | 0.997922 |

| Location | 3,199,877 – 3,199,968 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 78.68 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -29.80 |
| Energy contribution | -31.10 |
| Covariance contribution | 1.30 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.79 |
| SVM decision value | 2.96 |
| SVM RNA-class probability | 0.997922 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
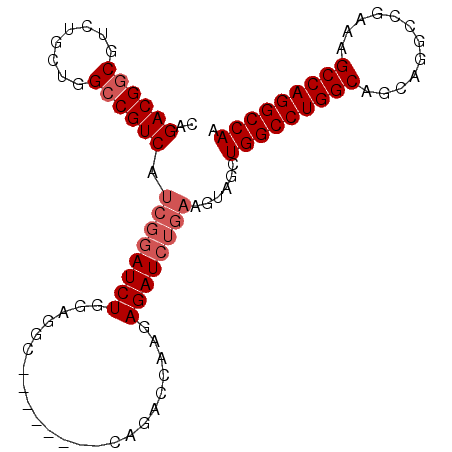
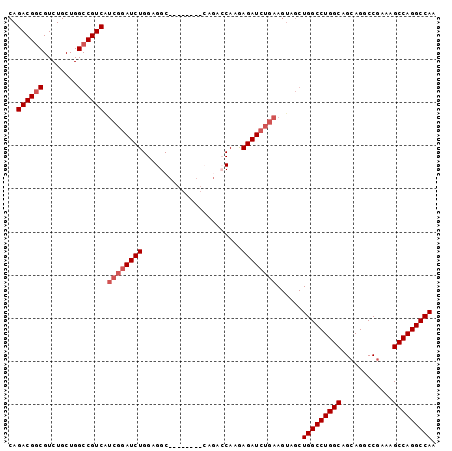
>2R_DroMel_CAF1 3199877 91 + 20766785 CAGACGGCGUCUGCUGGCCGUCAUCGGAUCUGGAGGC--------CAGACCAAGAGAUCUGAAGUAGCUGGCCUGGCAGCAGGCCGAAAGCCAGGCCAA ..((((((........)))))).((((((((...((.--------....))...))))))))......(((((((((............))))))))). ( -40.40) >DroSec_CAF1 6990 99 + 1 CAGACGGCGUCUGCUGGCCGUCAUCGGAUCUGGAGGCCUGGAGGCCAGACCAAGAGAUCUGAAGUAGCUGGCCUGGCAGCAGGCCGAAAGCCAGGCCAA ..((((((........)))))).((((((((((.((((....))))...))...))))))))......(((((((((............))))))))). ( -47.30) >DroSim_CAF1 10757 96 + 1 NNNNNNNNNNNNNNNNNNNNNNNNNNNAUCUGGAGGCC---AGACCAGACCAAGAGAUCUGAAGUAGCUGGCCUGGCAGCAGGCCGAAAGCCAGGCCAA ............................(((((...))---))).((((.(....).)))).......(((((((((............))))))))). ( -25.50) >DroEre_CAF1 7108 86 + 1 CAGACGGCGUCUGCUGGCCGUCAUCGGAUCUGGAGGC-------------CAAAAGAUCUGAAGUAGCUGGCCUGGCAGCAGGCCGAUAGCCAGGCCAA ..((((((........)))))).((((((((......-------------....))))))))......(((((((((............))))))))). ( -39.20) >DroYak_CAF1 7260 86 + 1 CAGACGUCGUCUGCUGGCCGUCGUCGGAUCUGGAGGC-------------CAAAAGAUCUGAAGUAGCUGGCCUGGCAGAAGGCCGAUAGCCAGGCCAA .........(((((((((((.(.((((((((......-------------....))))))))....).))))).)))))).((((........)))).. ( -36.00) >consensus CAGACGGCGUCUGCUGGCCGUCAUCGGAUCUGGAGGC________CAGACCAAGAGAUCUGAAGUAGCUGGCCUGGCAGCAGGCCGAAAGCCAGGCCAA ..((((((........)))))).((((((((.......................))))))))......(((((((((............))))))))). (-29.80 = -31.10 + 1.30)

| Location | 3,199,877 – 3,199,968 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 78.68 |
| Mean single sequence MFE | -36.24 |
| Consensus MFE | -27.46 |
| Energy contribution | -29.66 |
| Covariance contribution | 2.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.76 |
| SVM decision value | 2.86 |
| SVM RNA-class probability | 0.997457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
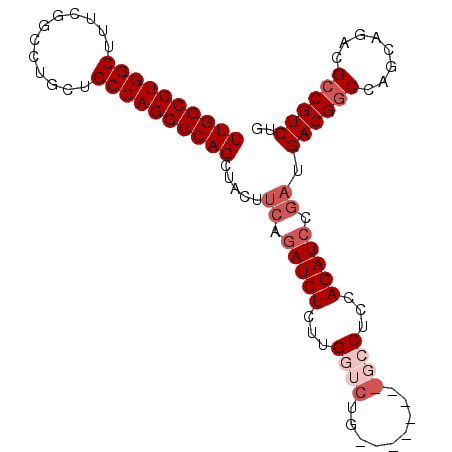
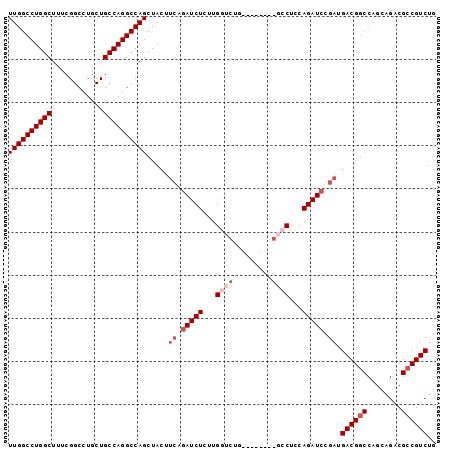
>2R_DroMel_CAF1 3199877 91 - 20766785 UUGGCCUGGCUUUCGGCCUGCUGCCAGGCCAGCUACUUCAGAUCUCUUGGUCUG--------GCCUCCAGAUCCGAUGACGGCCAGCAGACGCCGUCUG ((((((((((............))))))))))..............((((((((--------(...))))).)))).((((((........)))))).. ( -37.90) >DroSec_CAF1 6990 99 - 1 UUGGCCUGGCUUUCGGCCUGCUGCCAGGCCAGCUACUUCAGAUCUCUUGGUCUGGCCUCCAGGCCUCCAGAUCCGAUGACGGCCAGCAGACGCCGUCUG ((((((((((............)))))))))).....((.(((((...(((((((...)))))))...))))).)).((((((........)))))).. ( -46.50) >DroSim_CAF1 10757 96 - 1 UUGGCCUGGCUUUCGGCCUGCUGCCAGGCCAGCUACUUCAGAUCUCUUGGUCUGGUCU---GGCCUCCAGAUNNNNNNNNNNNNNNNNNNNNNNNNNNN .(((((((((............)))))))))((((..(((((((....)))))))..)---)))................................... ( -32.30) >DroEre_CAF1 7108 86 - 1 UUGGCCUGGCUAUCGGCCUGCUGCCAGGCCAGCUACUUCAGAUCUUUUG-------------GCCUCCAGAUCCGAUGACGGCCAGCAGACGCCGUCUG ((((((((((............)))))))))).....((.(((((...(-------------(...))))))).)).((((((........)))))).. ( -33.30) >DroYak_CAF1 7260 86 - 1 UUGGCCUGGCUAUCGGCCUUCUGCCAGGCCAGCUACUUCAGAUCUUUUG-------------GCCUCCAGAUCCGACGACGGCCAGCAGACGACGUCUG ((((((((((............))))))))))......(((((((.(((-------------(((((..........)).)))))).))).....)))) ( -31.20) >consensus UUGGCCUGGCUUUCGGCCUGCUGCCAGGCCAGCUACUUCAGAUCUCUUGGUCUG________GCCUCCAGAUCCGAUGACGGCCAGCAGACGCCGUCUG ((((((((((............)))))))))).....((.(((((...((((.........))))...))))).)).((((((........)))))).. (-27.46 = -29.66 + 2.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:43:23 2006