
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,753,943 – 19,754,082 |
| Length | 139 |
| Max. P | 0.779710 |

| Location | 19,753,943 – 19,754,042 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.02 |
| Mean single sequence MFE | -21.03 |
| Consensus MFE | -17.57 |
| Energy contribution | -17.82 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
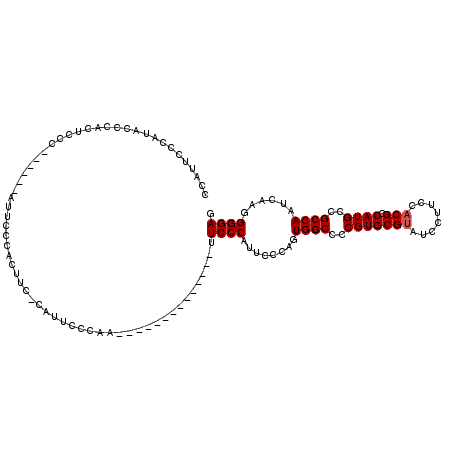
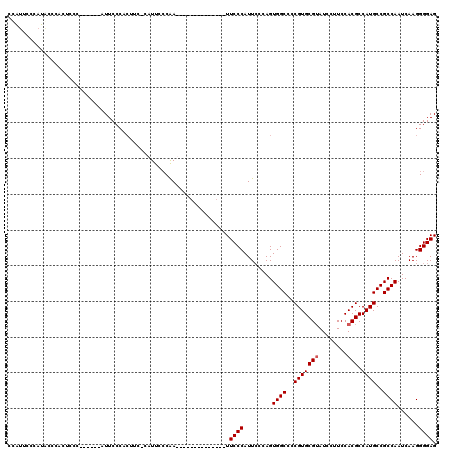
>2R_DroMel_CAF1 19753943 99 + 20766785 CCAUACCCAUACCAAUUCCC------AUUCCCACUUC-CAUUCCCGA--------------UUCCCAUUCCCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAG ....................------...........-.........--------------.((((........((((..(((((((........))).))))..))))......)))). ( -18.44) >DroSec_CAF1 6730 100 + 1 CCGUUCCCAUUCCCACUUCC------AUUCCCAAUUCCCAUUCCCAA--------------UUCCCAUUCUCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAG ....................------.....................--------------.((((........((((..(((((((........))).))))..))))......)))). ( -18.44) >DroEre_CAF1 6863 119 + 1 CUAUUUCCAUACCCAUUCCCACUUAUAUUCCCGCUUC-CAUUCGCAAUUCCGAAUUCCCAUUUCCCAUUCCCAGUGGCCCCGUGCGAAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAG ................((((.(((.............-.(((((......)))))..................(((((..((((.((.....))))))....))))).....))))))). ( -21.80) >DroYak_CAF1 9628 105 + 1 CCACUGCCAUGCCCACUGCC------AUUCCCACUUC-CAUUCCCAAUA-------CCCA-UUCCCAUUGGCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAG .........(.(((((((((------(..........-...........-------....-.......)))))))(((..(((((((........))).))))..)))......))).). ( -25.45) >consensus CCAUUCCCAUACCCACUCCC______AUUCCCACUUC_CAUUCCCAA______________UUCCCAUUCCCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAG ..............................................................((((........((((..(((((((........))).))))..))))......)))). (-17.57 = -17.82 + 0.25)

| Location | 19,753,976 – 19,754,082 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -25.17 |
| Consensus MFE | -21.92 |
| Energy contribution | -22.16 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748835 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
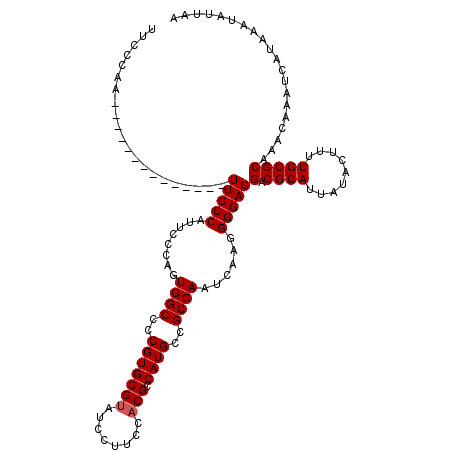
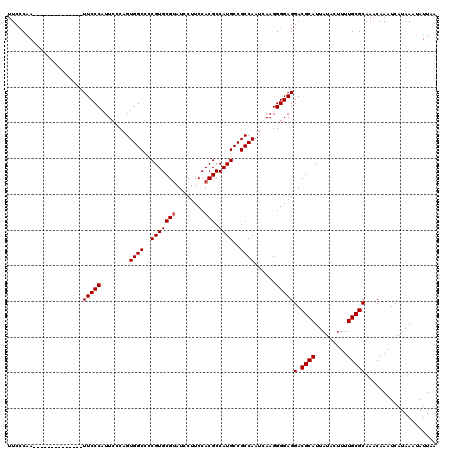
>2R_DroMel_CAF1 19753976 106 + 20766785 UUCCCGA--------------UUCCCAUUCCCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAGGACGCAUUAUACUUUUGCGCAAACAAAUCAUAAAUAUUAA .....((--------------((.((.(((((..((((..(((((((........))).))))..)))).....))))))).((((.........))))......))))........... ( -25.60) >DroSec_CAF1 6764 106 + 1 UUCCCAA--------------UUCCCAUUCUCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAGGACGCAUUAUACUUUUGCGCAAACAAAUCAUAAAUAUUAA .(((...--------------.((((........((((..(((((((........))).))))..))))......)))))))((((.........))))..................... ( -23.24) >DroEre_CAF1 6902 120 + 1 UUCGCAAUUCCGAAUUCCCAUUUCCCAUUCCCAGUGGCCCCGUGCGAAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAGGACGCAUUAUACUUUUGCGCAAACAAAUCAUAAAUAUUCA ...........(((((((...(((((.......(((((..((((.((.....))))))....))))).......))))))))((((.........))))................)))). ( -24.74) >DroYak_CAF1 9661 112 + 1 UUCCCAAUA-------CCCA-UUCCCAUUGGCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAGGACGCAUUAUACUUUUGCGCAAACAAAUCAUAAAUAUUAA .........-------.((.-(((((((((((.(((((...(((.(.......)))))))))...))))))...))))))).((((.........))))..................... ( -27.10) >consensus UUCCCAA______________UUCCCAUUCCCAGUGGCCCCGUGCGUAUCCUUCCACGCCAUGCCGCCAAUCAAGGGGAGGACGCAUUAUACUUUUGCGCAAACAAAUCAUAAAUAUUAA .....................(((((........((((..(((((((........))).))))..))))......)))))(.((((.........))))).................... (-21.92 = -22.16 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:07:51 2006