
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,684,290 – 19,684,397 |
| Length | 107 |
| Max. P | 0.508448 |

| Location | 19,684,290 – 19,684,397 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.16 |
| Mean single sequence MFE | -36.52 |
| Consensus MFE | -24.31 |
| Energy contribution | -24.98 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508448 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
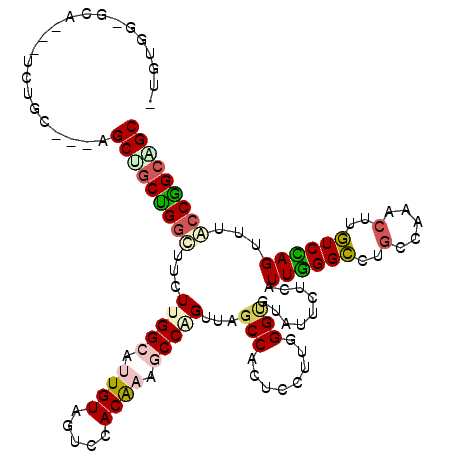
>2R_DroMel_CAF1 19684290 107 - 20766785 -------ACU---UCUUG---AGCUGCUGGCUUCUUGGCAUUGUAGUCGACAUAACCGGUCAGCCACUCCUUAGGUGUGUUUUCAUUGGGCCUUCCGAACUUGUCUAGCUUGCCGGCGGC -------...---.....---.(((((((((.....(((......)))((((....(((...(((........((((......)))).)))...)))....))))......))))))))) ( -32.30) >DroVir_CAF1 2049 114 - 1 CUGUGGUGCA---ACUGC---AGCCGCUGGCUUUUUGGCGUUGUAAUCCACAAAGCCUGUUAGCCAUUCCUUGGGCGUAUUCUCGUUGGGCCUGCCAAACUUGUCCAGUUUACCGGCAGC ..(((((((.---...))---).))))(((((....(((.((((.....)))).)))....)))))..(((..((((......))))))).((((((((((.....)))))...))))). ( -35.60) >DroGri_CAF1 1775 108 - 1 UUGUUGU--------UGC----GCUGCUGGCUUCUUGGCAUUGUAGUCCACAAAGCCAGUUAGCCACUCCUUGGGUGUAUUCUCAUUGGGCCUGCCAAACUUGUCCAGCUUGCCGGCAGC .......--------...----(((((((((...(((((.((((.....)))).)))))..(((((((.....)))).........(((((..(.....)..)))))))).))))))))) ( -36.50) >DroWil_CAF1 2042 117 - 1 CUGUGGCGUA---UCGGAAGCAGCUGCCGCCUUCUUGGCACUGUAGUCAACAAAACCAGUUAACCAUUCCUUGGGAGUAUUCUCAUUAGGCCUGCCAAACUUGUCUAGUUUACCGGCAGC ....(((.((---..((((((((.(((((......))))))))).((.(((.......))).))..)))).((((......)))).)).)))(((((((((.....)))))...)))).. ( -30.30) >DroMoj_CAF1 1728 114 - 1 UUGCGGGGCG---ACAGC---UGCGGCUGGCUUCUUGGCAUUGUAGUCCACAAAGCCUGUUAGCCAUUCUUUUGGUGUGUUUUCAUUGGGUCUGCCAAACUUAUCCAGUUUGCCAGCAGC ((((..((((---(..((---((.(((((((.....(((.((((.....)))).))).))))))).....(((((((..(((.....)))..)))))))......))))))))).)))). ( -36.60) >DroAna_CAF1 1641 113 - 1 -------GCUGCCUCUGCGGCGGCCACUGGUUUCUUGGCAUUGUAGUCCACGAAGCCAGUUAGCCACUCCUUUGGCGUGUUCUCGUUGGGCCUGCCAAACUUGUCCAGUUUACCGGCGGC -------((((((.(((.((((((.(((((((((.((((......).))).)))))))))..))).....(((((((.((((.....)))).)))))))...))))))......)))))) ( -47.80) >consensus _UGUGG_GCA___UCUGC___AGCUGCUGGCUUCUUGGCAUUGUAGUCCACAAAGCCAGUUAGCCACUCCUUGGGUGUAUUCUCAUUGGGCCUGCCAAACUUGUCCAGUUUACCGGCAGC ......................(((((((((...(((((.((((.....)))).)))))...(((........))).........((((((..(.....)..))))))...))))))))) (-24.31 = -24.98 + 0.67)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:07:07 2006