| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,569,112 – 19,569,223 |
| Length | 111 |
| Max. P | 0.940116 |

| Location | 19,569,112 – 19,569,223 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 65.79 |
| Mean single sequence MFE | -24.10 |
| Consensus MFE | -13.35 |
| Energy contribution | -14.35 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.940116 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 19569112 111 - 20766785 UCUGCACAU-U-GUUUUCUGCCUAAUAGGAAUUCAACUGUUAAACCGAACUAAG-AAAGAUUUUCCUAUUGCGUUGCUUAAUUGGCAAAUCUACUCUGCUGAAUUGAUGCAAUA .........-.-.......((..(((((((((((....(((......)))...)-)).....))))))))))(((((((((((((((.........)))).)))))).))))). ( -19.10) >DroSec_CAF1 796 87 - 1 UCUGCACUU-U-GUUUUGUGUCUAAUAGGAAUUCAAAAG-----------------------UUCCUAUUGCGUUGCUCAAU-GGCAAUGCUUCUCUGCGGAUUAGAUGCAGU- .(((((.((-(-(.(((((....(((((((((......)-----------------------))))))))((((((((....-))))))))......))))).))))))))).- ( -29.70) >DroSim_CAF1 1034 86 - 1 UCUGCACUU-U-GUUUUGUGACUAAUAGGAAUUCAACAG-----------------------CUCCUAUUGCGUUGCUCAAU-GGCAAUGCU-CUCUGCGGACUAGAUGCAGU- .(((((.((-(-(.(((((....(((((((.........-----------------------.)))))))((((((((....-)))))))).-....))))).))))))))).- ( -26.20) >DroYak_CAF1 1091 113 - 1 UUUGUGCUAUUGAGCUUAGGCCUAAAAAGAAUUCAACAGCUGAACCGAACUACUAAAAGAUUUUCCUGUUGCACUUCCAAAUUGGCAAUGCUCCUCCGUGGUUUCGAUGCAUC- ...((((.(((((((....(((......(((..((((((..(((.(............).)))..))))))...)))......)))...)).((.....))..))))))))).- ( -21.40) >consensus UCUGCACUU_U_GUUUUGUGCCUAAUAGGAAUUCAACAG_______________________UUCCUAUUGCGUUGCUCAAU_GGCAAUGCU_CUCUGCGGAUUAGAUGCAGU_ ((((((.................((((((((...............................))))))))((((((((.....)))))))).....))))))............ (-13.35 = -14.35 + 1.00)

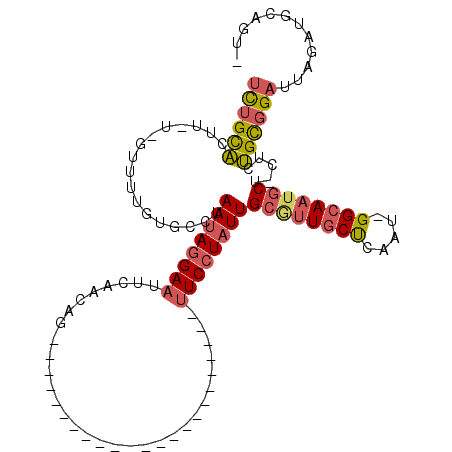
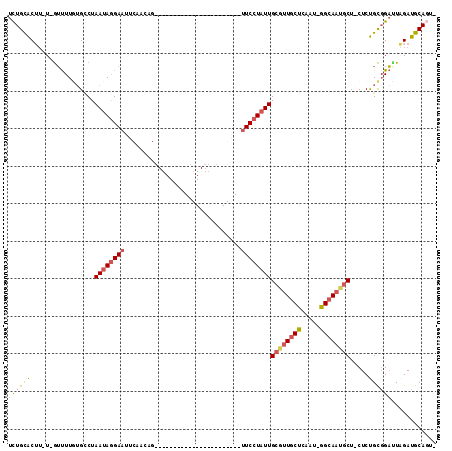
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:08 2006