| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,517,653 – 19,517,749 |
| Length | 96 |
| Max. P | 0.993921 |

| Location | 19,517,653 – 19,517,749 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 75.81 |
| Mean single sequence MFE | -36.32 |
| Consensus MFE | -18.14 |
| Energy contribution | -17.97 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.50 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 19517653 96 + 20766785 U---------GGAGACGGACUUGGAGGAC------------UGCUGCUUCUUGUACAGCUCUAUGGCCUCCUGCUCUAGCUUUAGCAGCUCCUGCUGCUCGUGCUGAAGUCGCUCCU .---------((((...((((((((((..------------.((((.........)))).......))))).....((((...((((((....))))))...))))))))).)))). ( -37.30) >DroPse_CAF1 14518 117 + 1 UAUGCUUUUCGGCAGUUGUCUUUGAAGAAUAUUGCCGGUCGUGCUGUUUCUUGUACAGCUCGAUGGACUCCUGCUCAAGCUUCAGCAGUUCCUGCUGCUCGAGCUGCAGCCGCUCUU ...((....(((((((..(((....)))..)))))))((((.(((((.......))))).))))((....((((...(((((.((((((....)))))).))))))))))))).... ( -44.10) >DroEre_CAF1 14401 96 + 1 U---------GGAGACGGACUUGGAAGAC------------UGCUGCUUCUUGUACAGCUCAAUGGCCUCCUGCUCCAGCUUUAGCAGCUCCUGCUGCUCGUGCUGAAGGCGCUCCU .---------((((.(((.((..((.(.(------------((.(((.....)))))))))...))))....(((.((((...((((((....))))))...))))..)))))))). ( -33.30) >DroYak_CAF1 14361 96 + 1 A---------GGAGAUGGACUUGGAAGAC------------UGCUGCUUCUUGUACAGCUCAAUGGCCUCCUGCUCUAGCUUCAGCAGCUCCUGCUGCUCGUGCUGAAGCCGCUCCU (---------((((..((.((..((.(.(------------((.(((.....)))))))))...))))....(((.((((...((((((....))))))...)))).)))..))))) ( -32.50) >DroAna_CAF1 15113 91 + 1 --------------ACGGAUUUCGAGGAC------------UGCUGCUUCUUGUACAGCUCUAUGGCUUCCUGUUCGAGCCUCAGCAGUUCCUGCUGUUCAAGCUGAAGGCGCUCUU --------------..(((..(((.((((------------((.(((.....)))))).))).)))..))).....((((..((((((...)))))).....((.....)))))).. ( -26.60) >DroPer_CAF1 14620 117 + 1 UAUGCUUUUCGGCAGUUGUCUUUGAAGAAUAUUGCCGGUCGUGCUGUUUCUUGUACAGCUCGAUGGACUCCUGCUCAAGCUUCAGCAGUUCCUGCUGCUCGAGCUGCAGCCGCUCUU ...((....(((((((..(((....)))..)))))))((((.(((((.......))))).))))((....((((...(((((.((((((....)))))).))))))))))))).... ( -44.10) >consensus U_________GGAGACGGACUUGGAAGAC____________UGCUGCUUCUUGUACAGCUCAAUGGCCUCCUGCUCAAGCUUCAGCAGCUCCUGCUGCUCGAGCUGAAGCCGCUCCU ..........................................((.(((((......(((.............)))..(((...((((((....))))))...)))))))).)).... (-18.14 = -17.97 + -0.17)
| Location | 19,517,653 – 19,517,749 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 75.81 |
| Mean single sequence MFE | -37.02 |
| Consensus MFE | -19.91 |
| Energy contribution | -20.38 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.54 |
| SVM decision value | 2.44 |
| SVM RNA-class probability | 0.993921 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
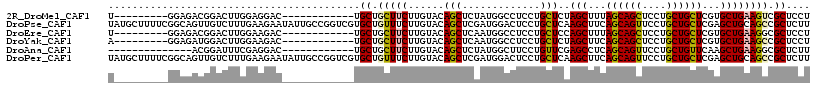
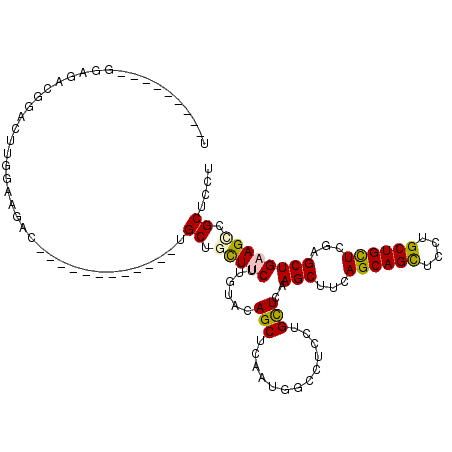
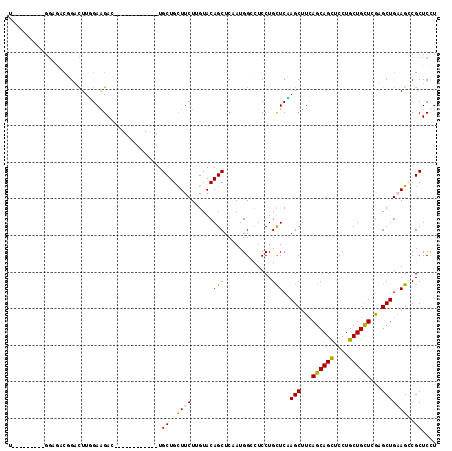
>2R_DroMel_CAF1 19517653 96 - 20766785 AGGAGCGACUUCAGCACGAGCAGCAGGAGCUGCUAAAGCUAGAGCAGGAGGCCAUAGAGCUGUACAAGAAGCAGCA------------GUCCUCCAAGUCCGUCUCC---------A .((((((.((..(((...((((((....))))))...)))..))..(((((.(.....(((((.......))))).------------).))))).....)).))))---------. ( -37.70) >DroPse_CAF1 14518 117 - 1 AAGAGCGGCUGCAGCUCGAGCAGCAGGAACUGCUGAAGCUUGAGCAGGAGUCCAUCGAGCUGUACAAGAAACAGCACGACCGGCAAUAUUCUUCAAAGACAACUGCCGAAAAGCAUA ....((((((.(.(((((((((((((...)))))...)))))))).).))))..(((.(((((.......))))).))).(((((....(((....)))....)))))....))... ( -42.90) >DroEre_CAF1 14401 96 - 1 AGGAGCGCCUUCAGCACGAGCAGCAGGAGCUGCUAAAGCUGGAGCAGGAGGCCAUUGAGCUGUACAAGAAGCAGCA------------GUCUUCCAAGUCCGUCUCC---------A .((((((.(((((((...((((((....))))))...)))))))..(((((.(.....(((((.......))))).------------).))))).....)).))))---------. ( -39.00) >DroYak_CAF1 14361 96 - 1 AGGAGCGGCUUCAGCACGAGCAGCAGGAGCUGCUGAAGCUAGAGCAGGAGGCCAUUGAGCUGUACAAGAAGCAGCA------------GUCUUCCAAGUCCAUCUCC---------U (((((..(((..(((...((((((....))))))...)))..))).(((((.(.....(((((.......))))).------------).)))))........))))---------) ( -34.90) >DroAna_CAF1 15113 91 - 1 AAGAGCGCCUUCAGCUUGAACAGCAGGAACUGCUGAGGCUCGAACAGGAAGCCAUAGAGCUGUACAAGAAGCAGCA------------GUCCUCGAAAUCCGU-------------- ....(((..(((.(((.....)))((((.(((((..(((((......).)))).....(((........)))))))------------))))).)))...)))-------------- ( -24.70) >DroPer_CAF1 14620 117 - 1 AAGAGCGGCUGCAGCUCGAGCAGCAGGAACUGCUGAAGCUUGAGCAGGAGUCCAUCGAGCUGUACAAGAAACAGCACGACCGGCAAUAUUCUUCAAAGACAACUGCCGAAAAGCAUA ....((((((.(.(((((((((((((...)))))...)))))))).).))))..(((.(((((.......))))).))).(((((....(((....)))....)))))....))... ( -42.90) >consensus AAGAGCGGCUUCAGCACGAGCAGCAGGAACUGCUGAAGCUAGAGCAGGAGGCCAUAGAGCUGUACAAGAAGCAGCA____________GUCUUCCAAGUCCAUCUCC_________A (((((.((((((.((.(.((((((((...)))))...))).).)).))))))).....(((((.......)))))..............))))........................ (-19.91 = -20.38 + 0.48)

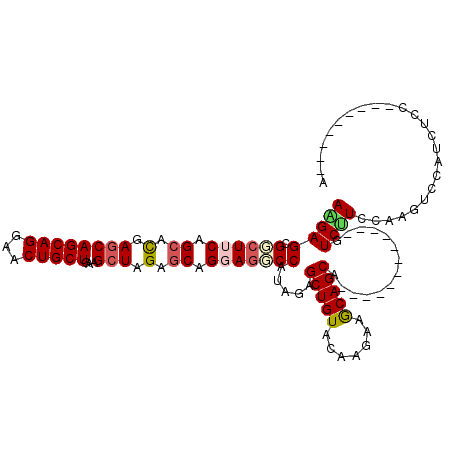
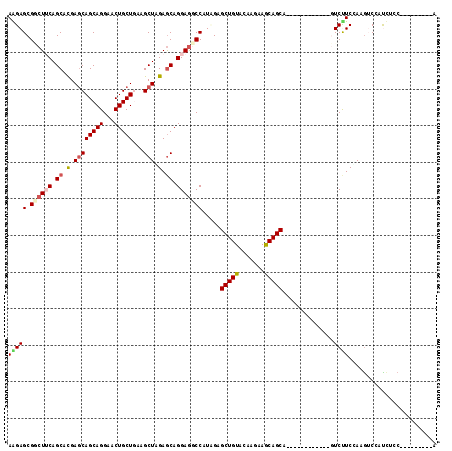
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:19 2006