
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,516,540 – 19,516,660 |
| Length | 120 |
| Max. P | 0.938273 |

| Location | 19,516,540 – 19,516,660 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.33 |
| Mean single sequence MFE | -38.82 |
| Consensus MFE | -25.80 |
| Energy contribution | -25.50 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.43 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938273 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
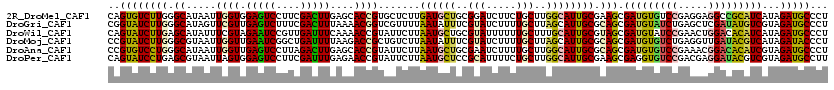
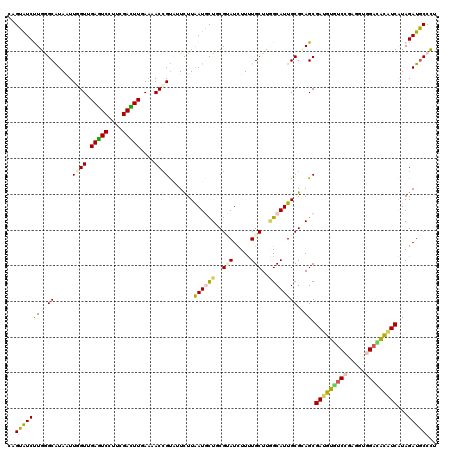
>2R_DroMel_CAF1 19516540 120 + 20766785 CAGUGUCUUGGGCAUAAUUGGUGGAGUCCUUCGACUUGAGCACCGUGCUCUUGAUGCUGCGGAUCUUCUGCUUGGCAUUGCGAAGCGAUGUGUCCGAGGAGGCCGCAUCAUAGAUGCCCU (((((((..((((((...((.(.(((((....))))).).))..))))))..))))))).((((((...((((.((...)).))))((((((.((.....)).))))))..)))).)).. ( -46.20) >DroGri_CAF1 18719 120 + 1 CGGUAUCUUGGGCAUAGUUCGUUGAGUCUUUCGACUUUAAAACGGUCGUUUUAAUAUUUCGUAUCUUUUGCUUAGCAUUGCGCAGCGAUGUAUCUGAGCUCGAUAUGUCGUAGAUGCCCU .........((((((.(..(((((((((....)))))...))))..).......(((..((((((....((((((((((((...))))))...))))))..))))))..))).)))))). ( -35.70) >DroWil_CAF1 14644 120 + 1 CAGUAUCUUGAGCAUAUUUCGUAGAAUCCGUUGAUUUCAAAACCGUAUUCUUAAUGCUGCGUAUUUUUUGCUUUGCAUUGCGUAGCGAUGUAUCCGAACUGGACACAUCAUAGAUGCCCU ..((((((((((.((((....(..((((....))))..).....)))).))))).((((((((.....(((...))).))))))))(((((.(((.....))).)))))...)))))... ( -30.70) >DroMoj_CAF1 17670 120 + 1 CCGUAUCUUGGGCGUAAUUGGUUGAAUCGGCUGAUUUUAAGACCGCUGUCUUAAUAUUUCGUAUCUUUUGCUUAGCAUUGCGCAGCGAUGUGUCUGAGGUUGAUACGUCAUAGAUACCCU ..((((((...(((((((((((...))))(((((..(((((((....)))))))......(((.....)))))))))))))))...((((((((.......))))))))..))))))... ( -33.50) >DroAna_CAF1 13947 120 + 1 CCGUGUCCUGGGCAUAAUUGGUUGAGUCCUUAGACUUGAGCACCGUAUUCUUAAUGCUGCGAAUCUUUUGCUUGGCAUUGCGCAGCGAUGUGUCCGAAACGGACACAUCGUAGAUGCCCU .........((((((...((.(..((((....))))..).)).........(((((((((((.....))))..)))))))....((((((((((((...))))))))))))..)))))). ( -46.50) >DroPer_CAF1 13603 120 + 1 CAGUAUCCUGAGCGUAAUUAGUGGAGUCCUUCGAUUUGAGAACCGUAUUCUUAAUGCUCCGCAUUUUCUGCUUGGCAUUGCGAAGCGAGGUGUCCGACGAGGAUACGUCGUAGAUGCCUU .........(((((.((...(((((((........(((((((.....))))))).)))))))...)).)))))((((((.....((((.((((((.....)))))).)))).)))))).. ( -40.30) >consensus CAGUAUCUUGGGCAUAAUUGGUUGAGUCCUUCGACUUGAAAACCGUAUUCUUAAUGCUGCGUAUCUUUUGCUUGGCAUUGCGCAGCGAUGUGUCCGAGGUGGACACAUCAUAGAUGCCCU ..((((((((.((.....((((.(((((....)))))....)))).......((((((..(((.....)))..)))))))).))).(((((((((.....)))))))))...)))))... (-25.80 = -25.50 + -0.30)

| Location | 19,516,540 – 19,516,660 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.33 |
| Mean single sequence MFE | -32.70 |
| Consensus MFE | -19.01 |
| Energy contribution | -19.65 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
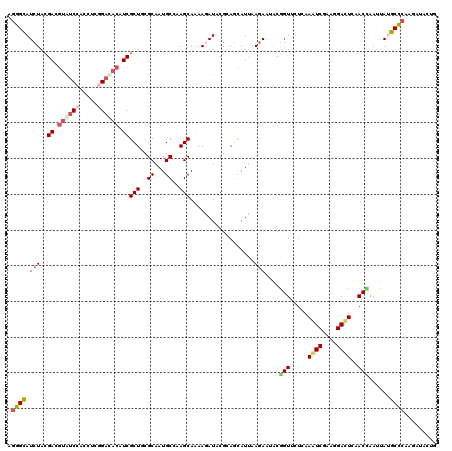
>2R_DroMel_CAF1 19516540 120 - 20766785 AGGGCAUCUAUGAUGCGGCCUCCUCGGACACAUCGCUUCGCAAUGCCAAGCAGAAGAUCCGCAGCAUCAAGAGCACGGUGCUCAAGUCGAAGGACUCCACCAAUUAUGCCCAAGACACUG .((((((((.((((((((...)).((((..(.((((((.((...)).)))).)).).))))..)))))))))....((((....((((....)))).)))).....)))))......... ( -38.90) >DroGri_CAF1 18719 120 - 1 AGGGCAUCUACGACAUAUCGAGCUCAGAUACAUCGCUGCGCAAUGCUAAGCAAAAGAUACGAAAUAUUAAAACGACCGUUUUAAAGUCGAAAGACUCAACGAACUAUGCCCAAGAUACCG .((((((........((((..((.(((........))).))..(((...)))...))))((.....(((((((....)))))))((((....))))...))....))))))......... ( -29.60) >DroWil_CAF1 14644 120 - 1 AGGGCAUCUAUGAUGUGUCCAGUUCGGAUACAUCGCUACGCAAUGCAAAGCAAAAAAUACGCAGCAUUAAGAAUACGGUUUUGAAAUCAACGGAUUCUACGAAAUAUGCUCAAGAUACUG .((((((....(((((((((.....)))))))))....((.(((((...((.........)).))))).(((((.(.(((........))).)))))).))....))))))......... ( -29.50) >DroMoj_CAF1 17670 120 - 1 AGGGUAUCUAUGACGUAUCAACCUCAGACACAUCGCUGCGCAAUGCUAAGCAAAAGAUACGAAAUAUUAAGACAGCGGUCUUAAAAUCAGCCGAUUCAACCAAUUACGCCCAAGAUACGG .((((........((((((.......((....))(((..((...))..)))....)))))).....(((((((....))))))).......................))))......... ( -23.40) >DroAna_CAF1 13947 120 - 1 AGGGCAUCUACGAUGUGUCCGUUUCGGACACAUCGCUGCGCAAUGCCAAGCAAAAGAUUCGCAGCAUUAAGAAUACGGUGCUCAAGUCUAAGGACUCAACCAAUUAUGCCCAGGACACGG .((((((((..((((((((((...))))))))))((((((.(((............)))))))))....)))....(((.....((((....))))..))).....)))))......... ( -43.70) >DroPer_CAF1 13603 120 - 1 AAGGCAUCUACGACGUAUCCUCGUCGGACACCUCGCUUCGCAAUGCCAAGCAGAAAAUGCGGAGCAUUAAGAAUACGGUUCUCAAAUCGAAGGACUCCACUAAUUACGCUCAGGAUACUG ..(((......((((......))))(((..(((.((((((((.(((...))).....))))))))..........(((((....))))).)))..))).........)))(((....))) ( -31.10) >consensus AGGGCAUCUACGACGUAUCCACCUCGGACACAUCGCUGCGCAAUGCCAAGCAAAAGAUACGCAGCAUUAAGAAUACGGUUCUCAAAUCGAAGGACUCAACCAAUUAUGCCCAAGAUACUG .((((.(((..((.((((((.....)))))).))(((..((...))..)))..................)))....(((.....((((....))))..)))......))))......... (-19.01 = -19.65 + 0.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:13 2006