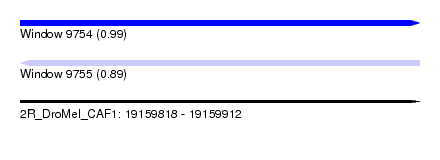

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,159,818 – 19,159,912 |
| Length | 94 |
| Max. P | 0.987221 |

| Location | 19,159,818 – 19,159,912 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 76.37 |
| Mean single sequence MFE | -25.98 |
| Consensus MFE | -14.19 |
| Energy contribution | -14.12 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.987221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
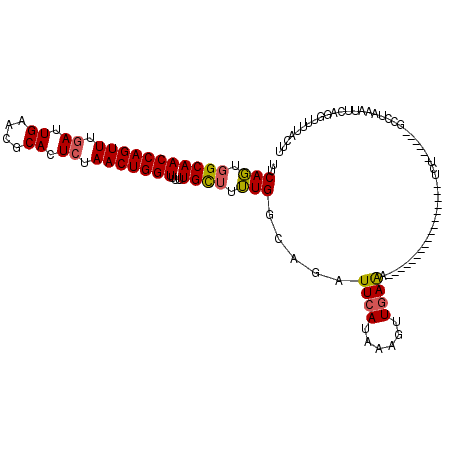
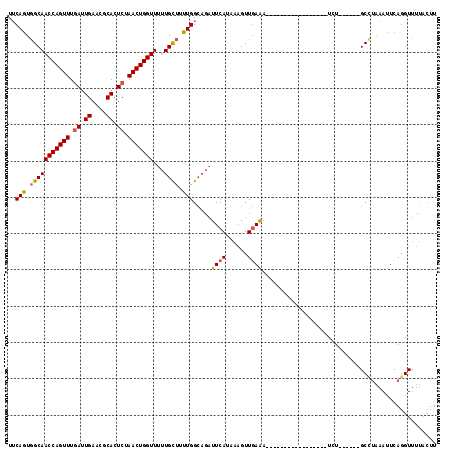
>2R_DroMel_CAF1 19159818 94 + 20766785 UACAGUGGCAACCAGUUUAAUUGAACGCACUCCAACUGGUUUUUGCU-UUGGCAGAUUCAUAAUGUUGAAA-----------------UCU------GCCUAAAUUCAGGUUUUACUU ...((..(.((((((((....((....))....)))))))).)..))-..((((((((((......)).))-----------------)))------))).................. ( -24.90) >DroSec_CAF1 43581 95 + 1 UUCAGUGGCAACCAGUUUGAUUGAACGCACUCUAACUGGUUUUUGUUUUUGGCAGAUUCAUUAAGUUGAAA-----------------UCU------GCUUAAAUUCAGGUUUUACUU ...(((((.((((((((.((.((....)).)).))))))))((((..(((((((((((((......)).))-----------------)))------)).))))..))))..))))). ( -25.30) >DroSim_CAF1 44626 95 + 1 UUCAGUGGCAACCAGUUUGAUUGAACGCACUCUAACUGGUUUUUGUUUUUGGCAGAUUCAUAAAGUUGAAA-----------------UCU------GCCUAAAUUCAGGUUUUUGUU ..(((..((((((((((.((.((....)).)).)))))))).(((..(((((((((((((......)).))-----------------)))------))).)))..)))))..))).. ( -28.30) >DroYak_CAF1 50501 114 + 1 UUCAUUAGCAACCAGUUUGAUUGAACGCACUCUAACUGGUUUAUGCCUAUGG----UUCAUUGAGUUCAGAAGGCAAUAAGUAUUUUUUUUGCCACAGCGAUAAUUCAGGUUUCACUU .((((..((((((((((.((.((....)).)).)))))))...)))..))))----....(((((((..(..(((((.(((....))).)))))....)...)))))))......... ( -25.40) >consensus UUCAGUGGCAACCAGUUUGAUUGAACGCACUCUAACUGGUUUUUGCUUUUGGCAGAUUCAUAAAGUUGAAA_________________UCU______GCCUAAAUUCAGGUUUUACUU ..(((.(((((((((((.((.((....)).)).)))))))...)))).))).....((((......))))................................................ (-14.19 = -14.12 + -0.06)

| Location | 19,159,818 – 19,159,912 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 76.37 |
| Mean single sequence MFE | -22.85 |
| Consensus MFE | -10.56 |
| Energy contribution | -10.38 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.46 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.885710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
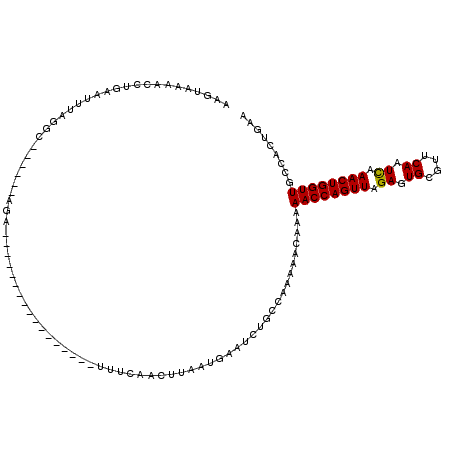
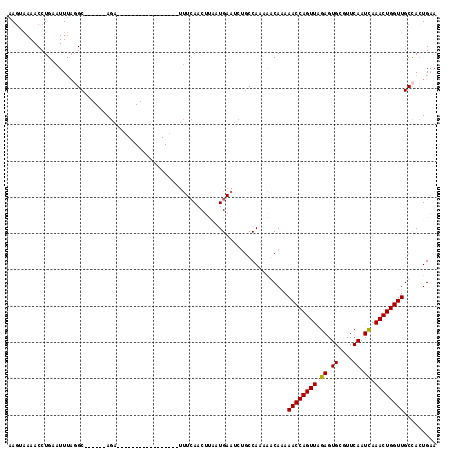
>2R_DroMel_CAF1 19159818 94 - 20766785 AAGUAAAACCUGAAUUUAGGC------AGA-----------------UUUCAACAUUAUGAAUCUGCCAA-AGCAAAAACCAGUUGGAGUGCGUUCAAUUAAACUGGUUGCCACUGUA .(((..............(((------(((-----------------((.((......))))))))))..-.(((...(((((((..(((.......))).)))))))))).)))... ( -24.20) >DroSec_CAF1 43581 95 - 1 AAGUAAAACCUGAAUUUAAGC------AGA-----------------UUUCAACUUAAUGAAUCUGCCAAAAACAAAAACCAGUUAGAGUGCGUUCAAUCAAACUGGUUGCCACUGAA .(((...............((------(((-----------------((.((......)))))))))..........((((((((.((.((....)).)).))))))))...)))... ( -20.10) >DroSim_CAF1 44626 95 - 1 AACAAAAACCUGAAUUUAGGC------AGA-----------------UUUCAACUUUAUGAAUCUGCCAAAAACAAAAACCAGUUAGAGUGCGUUCAAUCAAACUGGUUGCCACUGAA ..................(((------(((-----------------((.((......)))))))))).........((((((((.((.((....)).)).))))))))......... ( -23.60) >DroYak_CAF1 50501 114 - 1 AAGUGAAACCUGAAUUAUCGCUGUGGCAAAAAAAAUACUUAUUGCCUUCUGAACUCAAUGAA----CCAUAGGCAUAAACCAGUUAGAGUGCGUUCAAUCAAACUGGUUGCUAAUGAA .(((((...........)))))..(((((.((......)).)))))................----.(((.((((...(((((((.((.((....)).)).))))))))))).))).. ( -23.50) >consensus AAGUAAAACCUGAAUUUAGGC______AGA_________________UUUCAACUUAAUGAAUCUGCCAAAAACAAAAACCAGUUAGAGUGCGUUCAAUCAAACUGGUUGCCACUGAA .............................................................................((((((((.((.((....)).)).))))))))......... (-10.56 = -10.38 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:02:29 2006