
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,099,975 – 19,100,070 |
| Length | 95 |
| Max. P | 0.879592 |

| Location | 19,099,975 – 19,100,070 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 71.99 |
| Mean single sequence MFE | -22.77 |
| Consensus MFE | -9.34 |
| Energy contribution | -10.87 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.41 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879592 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
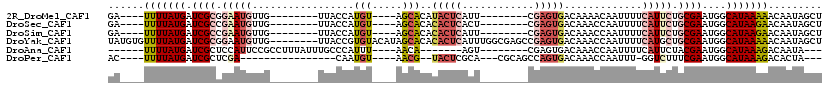
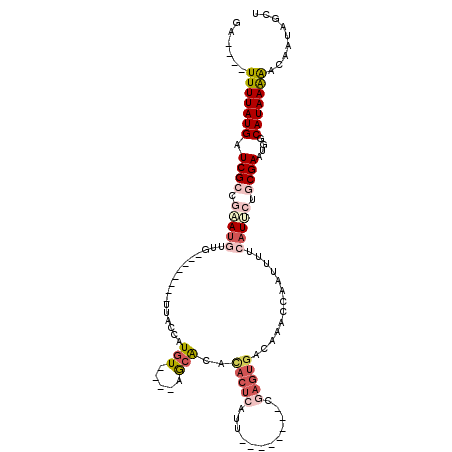
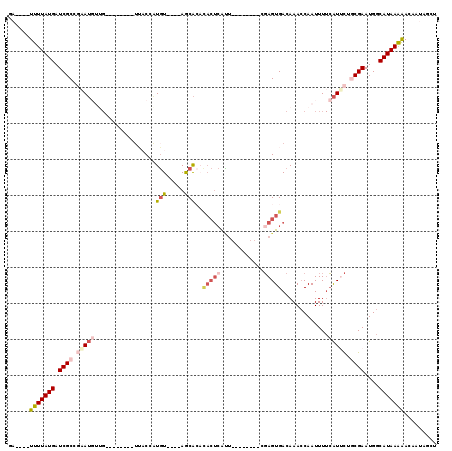
>2R_DroMel_CAF1 19099975 95 + 20766785 GA----UUUUAUGAUCGCGGAAUGUUG--------UUACCAUGU----AGCACAUACUCAUU--------CGAGUGACAAAACAAUUUUCAUUCUGCGAAUGGCAUAAAAACAAUAGCU ..----(((((((.((((((((((.((--------((((...))----))))..(((((...--------.))))).............))))))))))....)))))))......... ( -26.60) >DroSec_CAF1 42543 95 + 1 GA----UUUUAUGAUCGCCGAAUGUUG--------UUACCAUGU----AGCACACACUCACU--------CGAGUGACAAACCAAUUUUCAUUCUGCGAAUGGCAUAAGAACAAUAGCU ..----(((((((.((((.(((((.((--------((((...))----))))..(((((...--------.))))).............))))).))))....)))))))......... ( -24.90) >DroSim_CAF1 43409 95 + 1 GA----UUUUAUGAUCGCCGAAUGUUG--------UUACCAUGU----AGCACACACUCAUU--------CGAGUGACAAACCAAUUUUCAUUCUGCGAAUGGCAUAAGAACAAUAGCU ..----(((((((.((((.(((((.((--------((((...))----))))..(((((...--------.))))).............))))).))))....)))))))......... ( -24.90) >DroYak_CAF1 56392 111 + 1 UAUGUGUUUUAUGAUCGCGGAAUGUUG--------UUACCGUGUACAUAGCACACACUCAUUUGGCGAGCCGAGUGACAAACCAAUUUUCAUGCUGCGAAUGGCAUAAAAACAAUAGCU ..(((((.(((((..((((((((...)--------)).)))))..))))).))))).((((((((....))))))))........((((.(((((......))))).))))........ ( -29.80) >DroAna_CAF1 38323 91 + 1 ------UUUUAUGAUCGCUCCAUUCCGCCUUUAUUUGCCCAUUU----AACA-------AGU--------CGAGUGACAAACCAAUUUUCAUUCUACGAAUGGCAUAAAGACAAUA--- ------(((((((......((((((.((........))......----....-------.((--------.((((((.((.....)).)))))).)))))))))))))))......--- ( -14.60) >DroPer_CAF1 40376 86 + 1 AC----UUUUAUGAUCGCUCGA----------------CAAUGU----AACG--UACUCGCA---CGCAGCCAGUGACAAACCAAUUU-GGUCUUUCGAAUGGCAUAAAGACACUA--- ..----(((((((.(((.((((----------------......----...(--((((.((.---....)).))).))..(((.....-)))...)))).))))))))))......--- ( -15.80) >consensus GA____UUUUAUGAUCGCCGAAUGUUG________UUACCAUGU____AGCACACACUCAUU________CGAGUGACAAACCAAUUUUCAUUCUGCGAAUGGCAUAAAAACAAUAGCU ......(((((((.((((.(((((.................(((.....)))..(((((............))))).............))))).))))....)))))))......... ( -9.34 = -10.87 + 1.53)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:56 2006