
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 19,068,020 – 19,068,120 |
| Length | 100 |
| Max. P | 0.638326 |

| Location | 19,068,020 – 19,068,120 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 87.46 |
| Mean single sequence MFE | -28.40 |
| Consensus MFE | -20.09 |
| Energy contribution | -20.15 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.638326 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
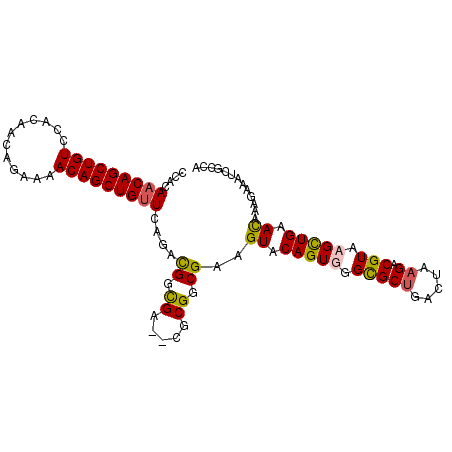
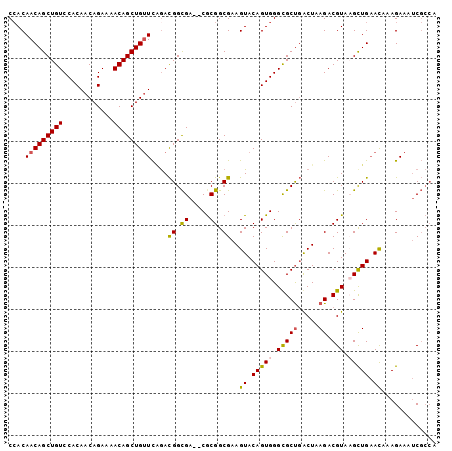
>2R_DroMel_CAF1 19068020 100 - 20766785 CCACAACAGCUGUCCACAACAGAAAACAGCUGUUCGAACGGCGA--CACGGCGAAGUACAGUGGGCGCUGGCUAAGACGUAAGCUGAACAAAGAAAUCGACA .....((((((((............))))))))((((.((.((.--..)).))..((.((((..(((((.....)).)))..)))).)).......)))).. ( -27.20) >DroSec_CAF1 10534 100 - 1 CCACAACAGCUGUCCACAACAGAAAACAGCUGUUCAGACGGCGA--CGCGGCGAAGUACAGUGGGUGCUGACUAAGACGUACGCUGAACAAAGAAAUCGCCA ........((((((...(((((.......)))))..))))))..--...(((((.((.(((((..((((.....)).))..))))).)).......))))). ( -29.20) >DroSim_CAF1 10417 100 - 1 CCACAACAGCUGUCCACAACAGAAAACAGCUGUUCAGACGGCGA--CGCGGCGAAGUACAGUGGGCGCUGACUAAGACGUACGCUGAACAAAGAAAUCGCCA ........((((((...(((((.......)))))..))))))..--...(((((.((.(((((..((((.....)).))..))))).)).......))))). ( -31.60) >DroYak_CAF1 23297 102 - 1 CAACAACAGCUGUCCACAACAGAAAACAGCUGAUUUGAUGGUGAAUCGCGGCGAAGUACAGUGGGCGCCCGCUUAGACGUAAGUUGCAUUAAGAAAACCCAA ......(((((((............)))))))....(((.....)))(((((...((..(((((....)))))...))....)))))............... ( -25.60) >consensus CCACAACAGCUGUCCACAACAGAAAACAGCUGUUCAGACGGCGA__CGCGGCGAAGUACAGUGGGCGCUGACUAAGACGUAAGCUGAACAAAGAAAUCGCCA ....(((((((((............)))))))))....((.((.....)).))..((.(((((.(((((.....)).))).))))).))............. (-20.09 = -20.15 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:39 2006