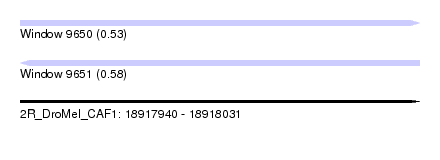
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,917,940 – 18,918,031 |
| Length | 91 |
| Max. P | 0.582338 |

| Location | 18,917,940 – 18,918,031 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 84.30 |
| Mean single sequence MFE | -24.80 |
| Consensus MFE | -17.50 |
| Energy contribution | -18.22 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.71 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.530110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18917940 91 + 20766785 ACGGCAAAUUACACAAAUUUUAUGUUGCCUACCUUUGGGCGCUCAAA---AGGCAACUUAAUAAGCACAAGUGUGUGCGGUUUCCCA-------GUACGUC ..((.((((..((((.((((......(((((....)))))(((...(---((....)))....)))..)))).))))..)))).)).-------....... ( -23.00) >DroSec_CAF1 17139 94 + 1 UCUUCAAAUUACACAAAUUUUAUGUUGCCUACCUUCGGGCGCUCAAAAGAAGGCAACUUAAUAAGCACAAGUGUGUGUGGUUUCCCA-------GUACGUC .....(((((((((.....(((.(((((((.((....))...((....))))))))).)))...(((....))))))))))))....-------....... ( -22.60) >DroSim_CAF1 20203 94 + 1 UCGGCAAAUUACACAAAUUUUAUGUUGCCUACCUUCGGGCGCUCAAAAGAAGGCAACUUAAUAAGCACAAGUGUGUGUGGUUUCCCA-------GUACGUC ..((.(((((((((.....(((.(((((((.((....))...((....))))))))).)))...(((....)))))))))))).)).-------....... ( -26.90) >DroYak_CAF1 18384 93 + 1 -CAGCAAAUUACACAAAUUUUAUGUUGCCUGCCUUCGGGCGCUUAAAAGAAGGCAACUUAAUAAGCACAAGUGUGUGCGGUUUCUCU-------GCUCGUG -.((((.............(((.((((((((((....))).((....)).))))))).)))...(((((....)))))........)-------))).... ( -26.70) >DroAna_CAF1 17884 99 + 1 UCAACAAAUUGCUGAAAUUUUAUGCUGCCUACCUUUGGGCGUUCCAAAGAGGGCAACUUAAUAAGCA--AGAGUGUGCGGUUUCUCUUCUAUGGGUUCCUC ...(((..(((((......(((.(.(((((..((((((.....)))))).))))).).)))..))))--)...)))..((...(((......)))..)).. ( -24.80) >consensus UCAGCAAAUUACACAAAUUUUAUGUUGCCUACCUUCGGGCGCUCAAAAGAAGGCAACUUAAUAAGCACAAGUGUGUGCGGUUUCCCA_______GUACGUC .....((((((((((...(((.((((((((......)))).........(((....)))....)))).)))..)))))))))).................. (-17.50 = -18.22 + 0.72)

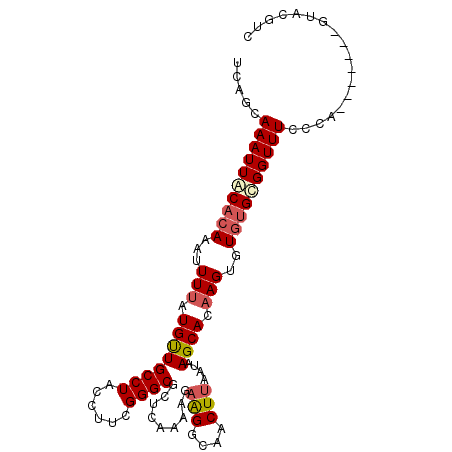
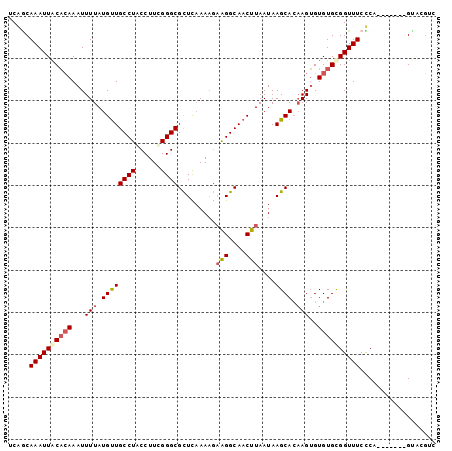
| Location | 18,917,940 – 18,918,031 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 84.30 |
| Mean single sequence MFE | -25.54 |
| Consensus MFE | -17.40 |
| Energy contribution | -18.72 |
| Covariance contribution | 1.32 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.582338 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
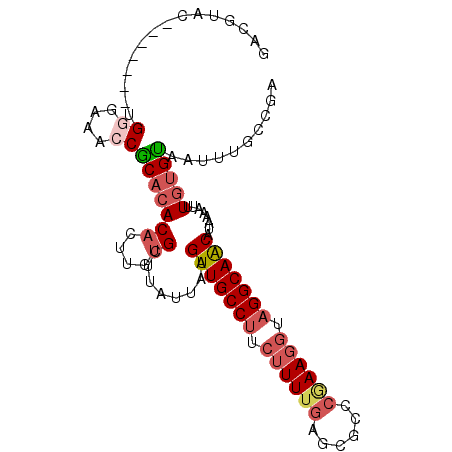
>2R_DroMel_CAF1 18917940 91 - 20766785 GACGUAC-------UGGGAAACCGCACACACUUGUGCUUAUUAAGUUGCCU---UUUGAGCGCCCAAAGGUAGGCAACAUAAAAUUUGUGUAAUUUGCCGU .......-------.((....))(((((....))))).........(((((---((.(.....).)))))))(((((((((.....)))))....)))).. ( -24.20) >DroSec_CAF1 17139 94 - 1 GACGUAC-------UGGGAAACCACACACACUUGUGCUUAUUAAGUUGCCUUCUUUUGAGCGCCCGAAGGUAGGCAACAUAAAAUUUGUGUAAUUUGAAGA ...((..-------(((....))).))((((..((..((((...(((((((.((((((......)))))).))))))))))).))..)))).......... ( -26.60) >DroSim_CAF1 20203 94 - 1 GACGUAC-------UGGGAAACCACACACACUUGUGCUUAUUAAGUUGCCUUCUUUUGAGCGCCCGAAGGUAGGCAACAUAAAAUUUGUGUAAUUUGCCGA ...((..-------(((....))).))((((..((..((((...(((((((.((((((......)))))).))))))))))).))..)))).......... ( -26.60) >DroYak_CAF1 18384 93 - 1 CACGAGC-------AGAGAAACCGCACACACUUGUGCUUAUUAAGUUGCCUUCUUUUAAGCGCCCGAAGGCAGGCAACAUAAAAUUUGUGUAAUUUGCUG- ....(((-------(((...((.(((((....))))).......(((((((.(((((........))))).))))))).........))....)))))).- ( -26.90) >DroAna_CAF1 17884 99 - 1 GAGGAACCCAUAGAAGAGAAACCGCACACUCU--UGCUUAUUAAGUUGCCCUCUUUGGAACGCCCAAAGGUAGGCAGCAUAAAAUUUCAGCAAUUUGUUGA .............(((((..........))))--).........((((((.(((((((.....)))))))..))))))........(((((.....))))) ( -23.40) >consensus GACGUAC_______UGGGAAACCGCACACACUUGUGCUUAUUAAGUUGCCUUCUUUUGAGCGCCCGAAGGUAGGCAACAUAAAAUUUGUGUAAUUUGCCGA ...............((....))(((((((....))........(((((((.((((((......)))))).)))))))........))))).......... (-17.40 = -18.72 + 1.32)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:00:32 2006