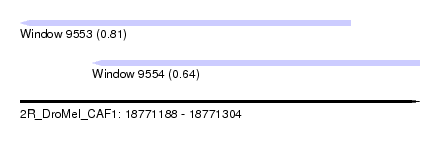

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,771,188 – 18,771,304 |
| Length | 116 |
| Max. P | 0.809775 |

| Location | 18,771,188 – 18,771,284 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 80.89 |
| Mean single sequence MFE | -25.02 |
| Consensus MFE | -21.10 |
| Energy contribution | -20.77 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.809775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
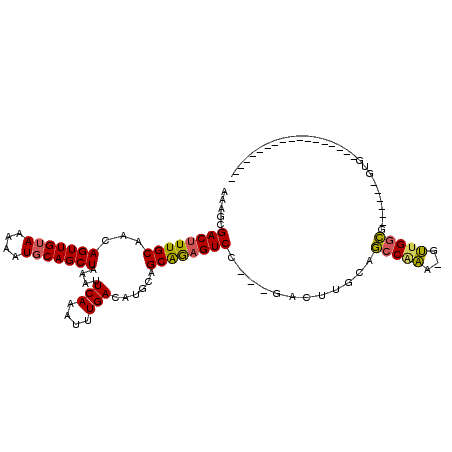
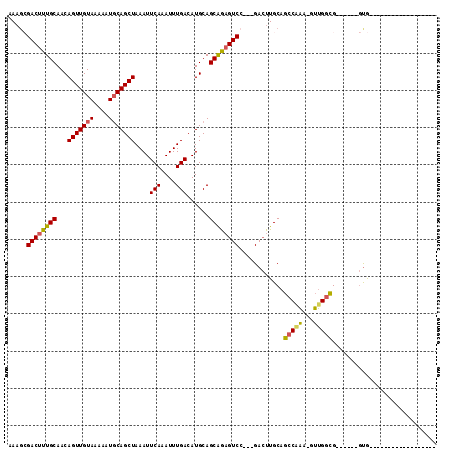
>2R_DroMel_CAF1 18771188 96 - 20766785 AAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAGCCAAA-GUUGGCGGUGGCGGUG------------------ ...((.(((.((((..(((((((....)))))))....(((....)))..))))(((.((((....))))))).((((..-..))))))).))....------------------ ( -27.00) >DroSec_CAF1 133717 90 - 1 AAAGCGACUUUGCAACAGUUGUAAAAAUCCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAGCCAAA-GUUGGCG------GUG------------------ ...(.((((((((...(((((........)))))....(((....)))......)))))))))........((.((((..-..)))).------)).------------------ ( -21.40) >DroSim_CAF1 136103 90 - 1 AAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAGCCAAA-GUUGGCG------GUG------------------ ...(.((((((((...(((((((....)))))))....(((....)))......)))))))))........((.((((..-..)))).------)).------------------ ( -24.20) >DroEre_CAF1 135668 84 - 1 AA---GACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCG---GACUUGCAGCCCAA-GUUGGCG------GUG------------------ ..---((((((((...(((((((....)))))))....(((....)))......)))))))).---.....((.(((...-...))).------)).------------------ ( -23.30) >DroYak_CAF1 136796 105 - 1 AAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGCGUCC---GACUUGCAGCCGAA-GUUGGCG------GUGGCGAUGAAGUUGGAGGUG .........(((((..(((((((....)))))))....(((....)))..)))))..((.(((---(((((...((((..-(....).------.))))....)))))))).)). ( -30.00) >DroAna_CAF1 134008 93 - 1 AAAGAGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCGAAGUCA---GAGUUGCCCGCGGAGGCUGAGG-CUGAGACA------------------ .....((((((((...(((((((....)))))))....(((....)))......)))))))).---..(((((((((....)).).))-)...))).------------------ ( -24.20) >consensus AAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCC___GACUUGCAGCCAAA_GUUGGCG______GUG__________________ .....((((((((...(((((((....)))))))....(((....)))......))))))))............(((((...)))))............................ (-21.10 = -20.77 + -0.33)

| Location | 18,771,209 – 18,771,304 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 80.89 |
| Mean single sequence MFE | -22.37 |
| Consensus MFE | -16.86 |
| Energy contribution | -16.92 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643047 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
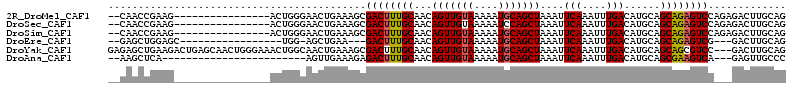
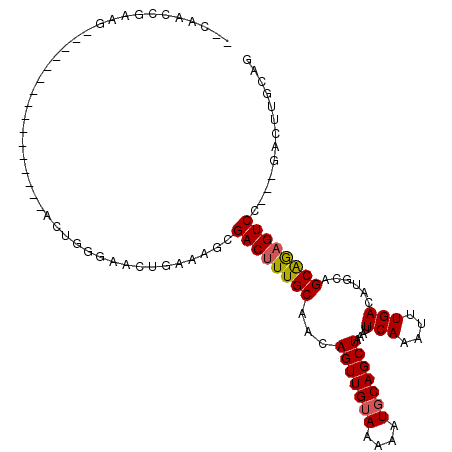
>2R_DroMel_CAF1 18771209 95 - 20766785 --CAACCGAAG----------------ACUGGGAACUGAAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAG --...(((...----------------..)))...(((.((((((((((((...(((((((....)))))))....(((....)))......))))))))....).))).))) ( -20.80) >DroSec_CAF1 133732 95 - 1 --CAACCGAAG----------------ACUGGGAACUGAAAGCGACUUUGCAACAGUUGUAAAAAUCCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAG --...(((...----------------..)))...(((.((((((((((((...(((((........)))))....(((....)))......))))))))....).))).))) ( -18.00) >DroSim_CAF1 136118 95 - 1 --CAACCGAAG----------------ACUGGGAACUGAAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCCAGAGACUUGCAG --...(((...----------------..)))...(((.((((((((((((...(((((((....)))))))....(((....)))......))))))))....).))).))) ( -20.80) >DroEre_CAF1 135683 87 - 1 --GAGCUGGAGC-----------------UGG-AGCUGAA---GACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCG---GACUUGCAG --..((.(((((-----------------...-.)))...---((((((((...(((((((....)))))))....(((....)))......)))))))).---..)).)).. ( -24.20) >DroYak_CAF1 136829 110 - 1 GAGAGCUGAAGACUGAGCAACUGGGAAACUGGCAACUGAAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGCGUCC---GACUUGCAG ................((((.(.(((..((((((((((...(((....)))..))))))).....((((..(((((....)))))...)))).)))..)))---.).)))).. ( -27.70) >DroAna_CAF1 134029 84 - 1 --AAGCUCA------------------------AGUUGAAAGAGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCGAAGUCA---GAGUUGCCC --.(((((.------------------------..((....))((((((((...(((((((....)))))))....(((....)))......)))))))).---))))).... ( -22.70) >consensus __CAACCGAAG________________ACUGGGAACUGAAAGCGACUUUGCAACAGUUGUAAAAAUGCAGCUAAAUUCAAAUUUGACAUGCAGCAGAGUCC___GACUUGCAG ...........................................((((((((...(((((((....)))))))....(((....)))......))))))))............. (-16.86 = -16.92 + 0.06)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:58 2006