
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,731,762 – 18,731,855 |
| Length | 93 |
| Max. P | 0.992234 |

| Location | 18,731,762 – 18,731,855 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 77.54 |
| Mean single sequence MFE | -23.16 |
| Consensus MFE | -14.40 |
| Energy contribution | -15.23 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924740 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
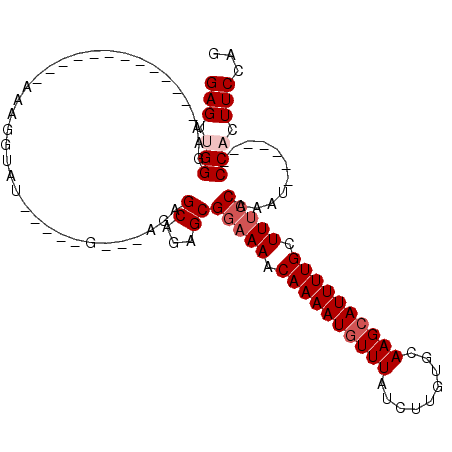
>2R_DroMel_CAF1 18731762 93 + 20766785 GAGUUGGUAAA---------AAAAAAGGUAUAAGCAG---AGCGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU-------CCACUUCCAG (((.(((....---------.................---.((.....))(((((.((((((((((.........)))))))))).)))))....-------))).)))... ( -19.80) >DroSec_CAF1 94622 84 + 1 GAGUUGGGAA-------------AAAGGUAU-----G---AGAGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU-------CCAAUUCCAG ((((((((..-------------........-----.---...((...))(((((.((((((((((.........)))))))))).)))))...)-------)))))))... ( -22.90) >DroSim_CAF1 95450 84 + 1 GAGUGGGGAA-------------AAAGCUAU-----G---AGAGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU-------CCACUUCCAG (((((((...-------------...(((.(-----(---....)).)))(((((.((((((((((.........)))))))))).)))))...)-------)))))).... ( -25.90) >DroEre_CAF1 95791 84 + 1 GAGUUGG-------------AGAAAAGGUAUA--------AGAGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU-------CCACUUCCAG (((.(((-------------(...........--------...((...))(((((.((((((((((.........)))))))))).)))))...)-------))).)))... ( -19.80) >DroYak_CAF1 96737 112 + 1 GAGUUGGGAAAAUAGGUAUGAAAAAAGGUAUGAUGAGAGUAGAGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAUCCACAAUCCACUUCCAG (..((((((..................((...((....))...)).....(((((.((((((((((.........)))))))))).)))))...))).)))..)........ ( -21.80) >DroAna_CAF1 93242 91 + 1 GAGGAGGGCCAGUGGAGGUG------GAAAU-----A---AAAGCUGGGCGACAAACAAAAUAUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU-------CCACUUCCAU .....(((..((((((..((------(((((-----(---((((((..((.((((..............)))))).))).))))).))))))..)-------)))))))).. ( -28.74) >consensus GAGUUGGGAA_____________AAAGGUAU_____G___AGAGCAGAGCGGAAAACAAAAUGUUUAUCUUGUGCAAGCAUUUUGCUUUCCAAAU_______CCACUUCCAG (((.(((....................................((...))(((((.((((((((((.........)))))))))).)))))...........))).)))... (-14.40 = -15.23 + 0.83)


| Location | 18,731,762 – 18,731,855 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 77.54 |
| Mean single sequence MFE | -24.25 |
| Consensus MFE | -15.51 |
| Energy contribution | -15.40 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.28 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992234 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
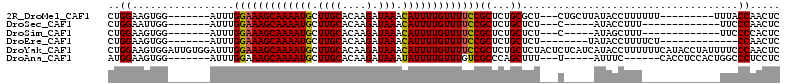
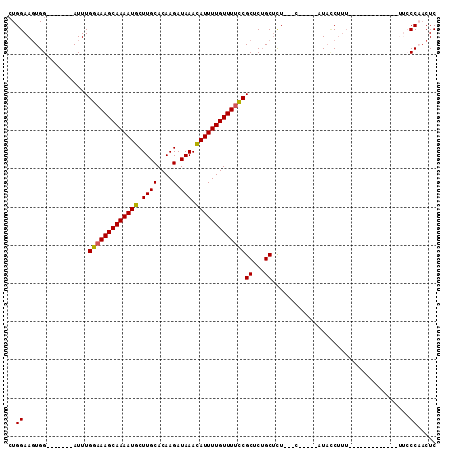
>2R_DroMel_CAF1 18731762 93 - 20766785 CUGGAAGUGG-------AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCGCU---CUGCUUAUACCUUUUUU---------UUUACCAACUC ..(((((..(-------(...(((((((((((((.((((....).))).)))))))))))))((.....)))---)..)))...))......---------........... ( -22.60) >DroSec_CAF1 94622 84 - 1 CUGGAAUUGG-------AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCUCU---C-----AUACCUUU-------------UUCCCAACUC ..((((..((-------....(((((((((((((.((((....).))).)))))))))))))((...))...---.-----...))...-------------))))...... ( -21.00) >DroSim_CAF1 95450 84 - 1 CUGGAAGUGG-------AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCUCU---C-----AUAGCUUU-------------UUCCCCACUC .....(((((-------....(((((((((((((.((((....).))).)))))))))))))(((.((....---)-----).)))...-------------....))))). ( -24.40) >DroEre_CAF1 95791 84 - 1 CUGGAAGUGG-------AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCUCU--------UAUACCUUUUCU-------------CCAACUC .((((((..(-------(..((((((((((((((.((((....).))).)))))))))))))).))..))..--------...........)-------------))).... ( -22.61) >DroYak_CAF1 96737 112 - 1 CUGGAAGUGGAUUGUGGAUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCUCUACUCUCAUCAUACCUUUUUUCAUACCUAUUUUCCCAACUC .((..((((((..(..((..((((((((((((((.((((....).))).)))))))))))))).))..)))))))..))................................. ( -28.40) >DroAna_CAF1 93242 91 - 1 AUGGAAGUGG-------AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAAUAUUUUGUUUGUCGCCCAGCUUU---U-----AUUUC------CACCUCCACUGGCCCUCCUC ..((.(((((-------(..((((((..((((.(((.((....(((((((.....)))))))))..))))))---)-----.))))------))..))))))..))...... ( -26.50) >consensus CUGGAAGUGG_______AUUUGGAAAGCAAAAUGCUUGCACAAGAUAAACAUUUUGUUUUCCGCUCUGCUCU___C_____AUACCUUU_____________UUCCCAACUC ..((.................(((((((((((((.((((....).))).)))))))))))))((...))....................................))..... (-15.51 = -15.40 + -0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:29 2006