| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,683,122 – 18,683,219 |
| Length | 97 |
| Max. P | 0.928060 |

| Location | 18,683,122 – 18,683,219 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 81.82 |
| Mean single sequence MFE | -24.42 |
| Consensus MFE | -13.76 |
| Energy contribution | -16.72 |
| Covariance contribution | 2.96 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.56 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.912528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
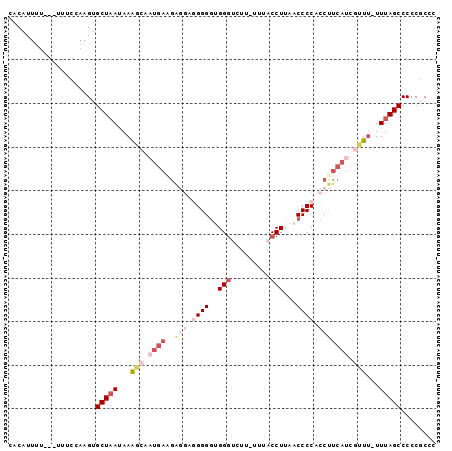
>2R_DroMel_CAF1 18683122 97 + 20766785 CACAUUUUUUUUUUCGAAGUGCUUAUAAAGCAAUGAAGAGGAGGGGCUGGGUCUU-UUUACCUUAACCCCACCUUCAUCGUUUUUUUAGCCCCCGCCC ..............((..(.(((...(((((.((((((.(..((((..((((...-...))))...)))).))))))).)))))...))))..))... ( -25.80) >DroSec_CAF1 46663 93 + 1 CACAUUUU---UUUCCAAGUGCUAAUAAAGCAAUGAGGAGGAGGGGGUGGGUCUU-UUUACCUUAACCCCACCUUCAUCGCUU-UUUAGCCCCCUCCC ........---..(((...((((.....))))....)))((((((((((((....-...........))))))......(((.-...))).)))))). ( -30.06) >DroSim_CAF1 47013 94 + 1 CACAUUUU---UUUCCAAGUGCUAAUAAAGCAAUGAGGAGGAGGGGGUGGGUCUUUUUUACCUUAACCCCACCUUCAUCGCUU-UUUAGCCCCCGCCC ........---.........(((((.(((((.((((((.(..((((..((((.......))))...)))).))))))).))))-))))))........ ( -29.20) >DroEre_CAF1 47629 88 + 1 CACAUUUU---UUUCCAAGUGCUAAUAAAGCGAUGAAGA----GGGGAGGCGCUU-UUUACCUUAACCCCACCCUCAGCAUU--UUUAGCCCCCGCCC ........---.........(((((......((((..((----(((..((.(.((-........)))))..)))))..))))--.)))))........ ( -16.30) >DroYak_CAF1 47049 79 + 1 CACAUU-----UUUCCAAGUGCUAAUAAG--------AG----GGGGCGGGGCUC-UCUACCUUAACCCCACCUUCAUAAUUU-UUUAGCCCCCGCCC ..((((-----(....)))))........--------.(----(((((((((...-..........)))).......(((...-.))))))))).... ( -20.72) >consensus CACAUUUU___UUUCCAAGUGCUAAUAAAGCAAUGAAGAGGAGGGGGUGGGUCUU_UUUACCUUAACCCCACCUUCAUCGUUU_UUUAGCCCCCGCCC ....................(((((...(((.((((..(((.((((..(((.........)))...)))).))))))).)))...)))))........ (-13.76 = -16.72 + 2.96)


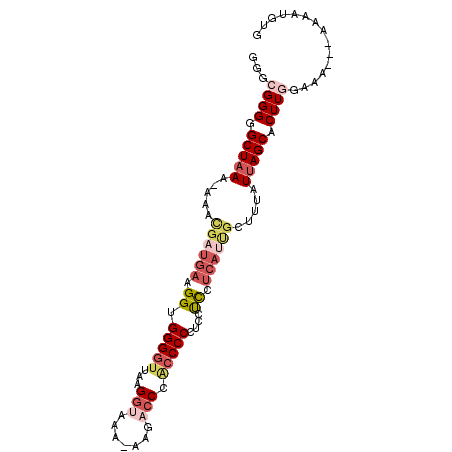
| Location | 18,683,122 – 18,683,219 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 81.82 |
| Mean single sequence MFE | -30.00 |
| Consensus MFE | -13.60 |
| Energy contribution | -15.92 |
| Covariance contribution | 2.32 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.45 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928060 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
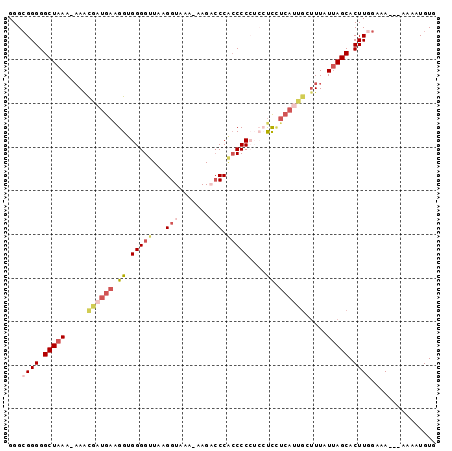
>2R_DroMel_CAF1 18683122 97 - 20766785 GGGCGGGGGCUAAAAAAACGAUGAAGGUGGGGUUAAGGUAAA-AAGACCCAGCCCCUCCUCUUCAUUGCUUUAUAAGCACUUCGAAAAAAAAAAUGUG ...((((((((.(.(((.(((((((((.((((((..(((...-...))).)))))).)))..)))))).))).).))).))))).............. ( -30.90) >DroSec_CAF1 46663 93 - 1 GGGAGGGGGCUAAA-AAGCGAUGAAGGUGGGGUUAAGGUAAA-AAGACCCACCCCCUCCUCCUCAUUGCUUUAUUAGCACUUGGAAA---AAAAUGUG ..(((...((((((-((((((((((((.((((....(((...-...)))...)))).)))..)))))))))).))))).))).....---........ ( -33.70) >DroSim_CAF1 47013 94 - 1 GGGCGGGGGCUAAA-AAGCGAUGAAGGUGGGGUUAAGGUAAAAAAGACCCACCCCCUCCUCCUCAUUGCUUUAUUAGCACUUGGAAA---AAAAUGUG ...((((.((((((-((((((((((((.((((....(((.......)))...)))).)))..)))))))))).))))).))))....---........ ( -34.80) >DroEre_CAF1 47629 88 - 1 GGGCGGGGGCUAAA--AAUGCUGAGGGUGGGGUUAAGGUAAA-AAGCGCCUCCCC----UCUUCAUCGCUUUAUUAGCACUUGGAAA---AAAAUGUG ...((((.(((((.--((.((((((((.((((...((((...-....))))))))----))))))..)).)).))))).))))....---........ ( -26.50) >DroYak_CAF1 47049 79 - 1 GGGCGGGGGCUAAA-AAAUUAUGAAGGUGGGGUUAAGGUAGA-GAGCCCCGCCCC----CU--------CUUAUUAGCACUUGGAAA-----AAUGUG (((.((((((....-..(((.....)))((((((........-.)))))))))))----).--------)))....(((.((....)-----).))). ( -24.10) >consensus GGGCGGGGGCUAAA_AAACGAUGAAGGUGGGGUUAAGGUAAA_AAGACCCACCCCCUCCUCCUCAUUGCUUUAUUAGCACUUGGAAA___AAAAUGUG ...((((.(((((.....((((((.((.(((((...(((.......))).)))))....)).)))))).....))))).))))............... (-13.60 = -15.92 + 2.32)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:15 2006