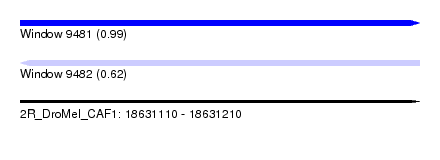

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,631,110 – 18,631,210 |
| Length | 100 |
| Max. P | 0.987868 |

| Location | 18,631,110 – 18,631,210 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 91.81 |
| Mean single sequence MFE | -36.76 |
| Consensus MFE | -28.68 |
| Energy contribution | -28.72 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.86 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.10 |
| SVM RNA-class probability | 0.987868 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18631110 100 + 20766785 UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGGUUGAAGGACACGAAAGUGGGCCACGGCAUAAGCCAGGGAAGCAGGGCAGGUUGACGACACACAAA ..((((((.(((..((((..((((.(.((.(((.((((..((.(((....)))..)).))))...))))).)..)))))))).))))).))))....... ( -38.50) >DroSec_CAF1 41738 97 + 1 UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGGUUGAAGGACACGAAAGUGGGCCAGGACAUAAGCCAGGGAAGCAGGGCAGGUUGG---CACACAAA ..((((.((.(((.(((((((....(((((.((......)).)))))..)))))))))).......(((..(....)..)))..)))))---)....... ( -38.00) >DroSim_CAF1 38140 96 + 1 UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGGUUGAAGGACACGAAAGUGGGCCAGGACAUAAGCCAGGG-AGCAGGGCAGGUUGG---CACACAAA ..((((.((.(((.(((((((....(((((.((......)).)))))..)))))))))).......(((.(..-..)..)))..)))))---)....... ( -35.70) >DroEre_CAF1 43900 92 + 1 UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGCUUGAAGGACACGAAAGUGGGCCAGGACAUAAGCCAGGCAAA-----CAGGUUGG---CACACAAA ..(((.....(((.(((((((....(((((.((......)).)))))..)))))))))))))....(((((.(...-----..).))))---)....... ( -36.20) >DroYak_CAF1 40676 92 + 1 UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGGCUGAAGGACACGAAAGUGGGCCAGGACCUAAGCCAGGGAAA-----CAGGUUGG---CACACAAA ..(((.....(((.(((((((....(((((.((......)).)))))..))))))))))......((((..(....-----).))))))---)....... ( -35.40) >consensus UAGUCGCACACUGAGCCCACUGCUACGUGUGCUGGUUGAAGGACACGAAAGUGGGCCAGGACAUAAGCCAGGGAAGCAGGGCAGGUUGG___CACACAAA ..(((.....(((.(((((((....(((((.((......)).)))))..)))))))))))))...((((..............))))............. (-28.68 = -28.72 + 0.04)

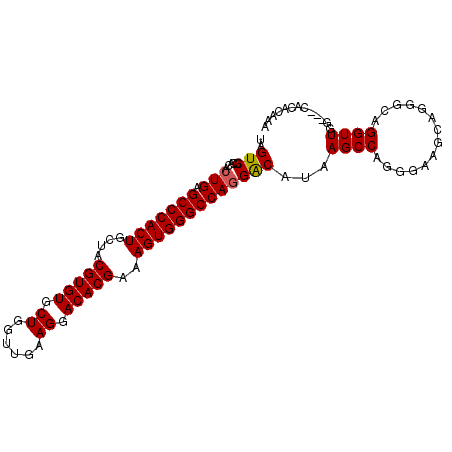
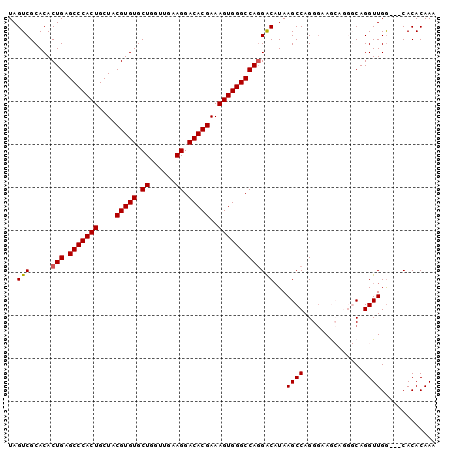
| Location | 18,631,110 – 18,631,210 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 91.81 |
| Mean single sequence MFE | -30.52 |
| Consensus MFE | -25.94 |
| Energy contribution | -25.98 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.621682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18631110 100 - 20766785 UUUGUGUGUCGUCAACCUGCCCUGCUUCCCUGGCUUAUGCCGUGGCCCACUUUCGUGUCCUUCAACCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA .......((((.((..(((............(((....)))..((((((((..(((((.((......)).)))))....)))))))))))..)))))).. ( -34.00) >DroSec_CAF1 41738 97 - 1 UUUGUGUG---CCAACCUGCCCUGCUUCCCUGGCUUAUGUCCUGGCCCACUUUCGUGUCCUUCAACCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA .(((..((---(((....((...)).....)))).......((((((((((..(((((.((......)).)))))....))))))).))).)..)))... ( -29.60) >DroSim_CAF1 38140 96 - 1 UUUGUGUG---CCAACCUGCCCUGCU-CCCUGGCUUAUGUCCUGGCCCACUUUCGUGUCCUUCAACCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA .(((..((---(((....((...)).-...)))).......((((((((((..(((((.((......)).)))))....))))))).))).)..)))... ( -30.20) >DroEre_CAF1 43900 92 - 1 UUUGUGUG---CCAACCUG-----UUUGCCUGGCUUAUGUCCUGGCCCACUUUCGUGUCCUUCAAGCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA .(((..((---(((..(..-----...)..)))).......((((((((((..(((((.((......)).)))))....))))))).))).)..)))... ( -29.50) >DroYak_CAF1 40676 92 - 1 UUUGUGUG---CCAACCUG-----UUUCCCUGGCUUAGGUCCUGGCCCACUUUCGUGUCCUUCAGCCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA ...((.((---((((((((-----...(....)..))))).((((((((((..(((((.((......)).)))))....))))))).))))).))))).. ( -29.30) >consensus UUUGUGUG___CCAACCUGCCCUGCUUCCCUGGCUUAUGUCCUGGCCCACUUUCGUGUCCUUCAACCAGCACACGUAGCAGUGGGCUCAGUGUGCGACUA .(((..(........................(((....)))((((((((((..(((((.((......)).)))))....))))))).))).)..)))... (-25.94 = -25.98 + 0.04)

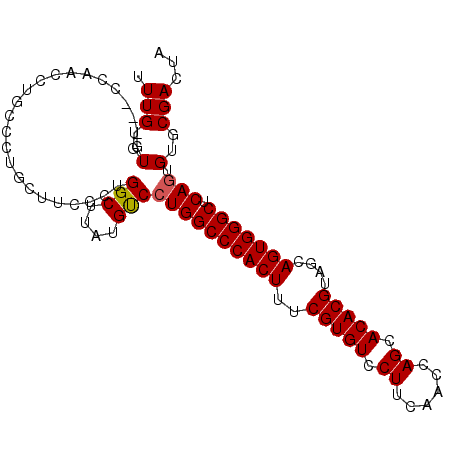
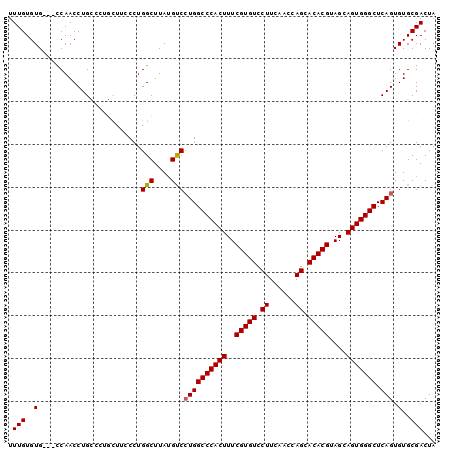
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:48 2006