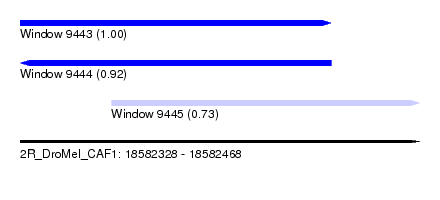
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,582,328 – 18,582,468 |
| Length | 140 |
| Max. P | 0.997920 |

| Location | 18,582,328 – 18,582,437 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 72.30 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -11.28 |
| Energy contribution | -10.96 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.31 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.41 |
| SVM decision value | 2.96 |
| SVM RNA-class probability | 0.997920 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18582328 109 + 20766785 UGAUAAUAUACAGAAGUACAAAGGCGAAAAUACCAAGAGAAUGAUGUUAGUGUGUGUGUUAGG-UGGGAAACUUGGCUGGGACUGUAGCUAAGUUCAACACCAGGCGGAG ...(((((((((.(...(((.(..(.............)..)..)))...).)))))))))((-((...((((((((((......))))))))))...))))........ ( -26.92) >DroSec_CAF1 7131 109 + 1 GGAUAAAAUACCGAAGUGCGCACGGAAAAAUACCAAGAGGAUGAUGUUAGUGUGGAUGUUUGG-UGGGAAACUUGGCUGGGGAUGUAGCUGAGUUCAACACCAGGCGGAG ..........((...((.(((((.((...((.((....))))....)).))))).))((((((-((...((((..((((......))))..))))...)))))))))).. ( -34.10) >DroSim_CAF1 4232 108 + 1 GGA-AAAAUACCGAAGUACGCACGGAAAAAUACCAAGAGGAUGAUGUUAGUGUGGAUGUUUGG-UGGGAAACUUGGUUGGGGAUGUAGCUGAGUUCAACACCAGGCGGAG ...-......((......(((((.((...((.((....))))....)).)))))...((((((-((...((((..((((......))))..))))...)))))))))).. ( -30.60) >DroEre_CAF1 7128 87 + 1 GGA-GAUAUACCGGAGAGCA-ACGGAAAAAUACCGAAAG----GUGU-----------AUGGGGUGGAAAACUUGGCUGAGGAUGUAGCAAAGAUCGAAACCAC------ ...-......(((.......-.)))....(((((....)----))))-----------.(((..(.((...(((.((((......)))).))).)).)..))).------ ( -22.20) >DroYak_CAF1 7276 95 + 1 GGA-UAAAUACCGAAGACCG-GCGGAAAAAUACCAAAAG----AUAUUAGAGUGGGUGUUGGGGUGGA-AACUUUGCUGAGGAUGUAGCAAGGUUCAAAACC-------- ...-......((((..(((.-((.....((((.(....)----.))))...)).))).))))(((.((-(.((((((((......)))))))))))...)))-------- ( -22.80) >consensus GGA_AAAAUACCGAAGUACG_ACGGAAAAAUACCAAGAG_AUGAUGUUAGUGUGGAUGUUGGG_UGGGAAACUUGGCUGGGGAUGUAGCUAAGUUCAACACCAGGCGGAG ((........)).................................................((.((...((((((((((......))))))))))...))))........ (-11.28 = -10.96 + -0.32)

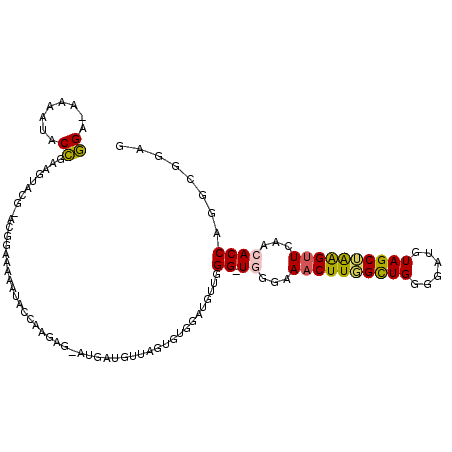
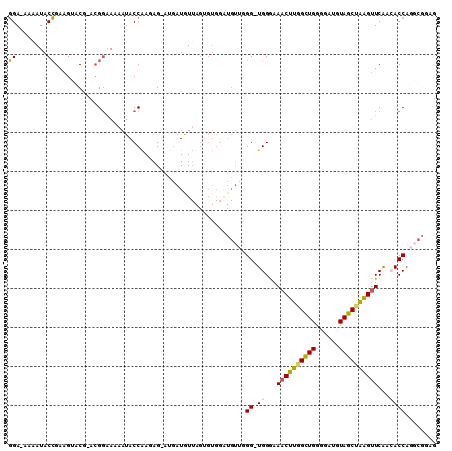
| Location | 18,582,328 – 18,582,437 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 72.30 |
| Mean single sequence MFE | -17.30 |
| Consensus MFE | -4.44 |
| Energy contribution | -6.04 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.26 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.917414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18582328 109 - 20766785 CUCCGCCUGGUGUUGAACUUAGCUACAGUCCCAGCCAAGUUUCCCA-CCUAACACACACACUAACAUCAUUCUCUUGGUAUUUUCGCCUUUGUACUUCUGUAUAUUAUCA ....((..((((..((((((.(((........))).))))))..))-))................((((......))))......))...(((((....)))))...... ( -19.50) >DroSec_CAF1 7131 109 - 1 CUCCGCCUGGUGUUGAACUCAGCUACAUCCCCAGCCAAGUUUCCCA-CCAAACAUCCACACUAACAUCAUCCUCUUGGUAUUUUUCCGUGCGCACUUCGGUAUUUUAUCC ....((.(((((..(((((..(((........)))..)))))..))-)))........(((.........((....)).........))).))((....))......... ( -20.97) >DroSim_CAF1 4232 108 - 1 CUCCGCCUGGUGUUGAACUCAGCUACAUCCCCAACCAAGUUUCCCA-CCAAACAUCCACACUAACAUCAUCCUCUUGGUAUUUUUCCGUGCGUACUUCGGUAUUUU-UCC ...(((.(((.((((....))))..........((((((.......-..........................))))))......))).)))(((....)))....-... ( -17.11) >DroEre_CAF1 7128 87 - 1 ------GUGGUUUCGAUCUUUGCUACAUCCUCAGCCAAGUUUUCCACCCCAU-----------ACAC----CUUUCGGUAUUUUUCCGU-UGCUCUCCGGUAUAUC-UCC ------((((.......(((.(((........))).)))....)))).....-----------....----.....(((((....(((.-.......))).)))))-... ( -13.10) >DroYak_CAF1 7276 95 - 1 --------GGUUUUGAACCUUGCUACAUCCUCAGCAAAGUU-UCCACCCCAACACCCACUCUAAUAU----CUUUUGGUAUUUUUCCGC-CGGUCUUCGGUAUUUA-UCC --------(((...((((.(((((........))))).)))-)..)))..............(((((----(....)))))).....((-((.....)))).....-... ( -15.80) >consensus CUCCGCCUGGUGUUGAACUUAGCUACAUCCCCAGCCAAGUUUCCCA_CCAAACACCCACACUAACAUCAU_CUCUUGGUAUUUUUCCGU_CGCACUUCGGUAUUUU_UCC ........((((..((((((.(((........))).))))))..)).))............................................................. ( -4.44 = -6.04 + 1.60)

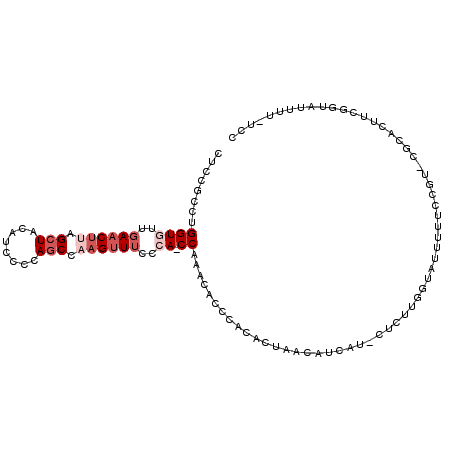
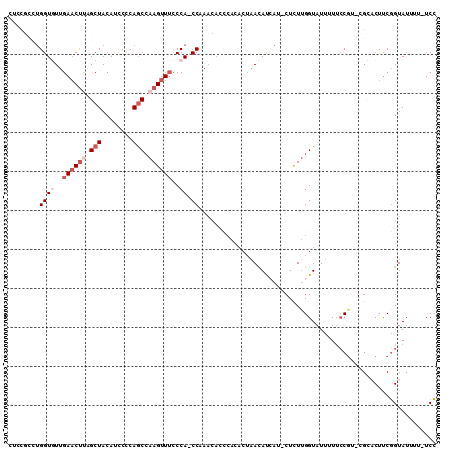
| Location | 18,582,360 – 18,582,468 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 64.26 |
| Mean single sequence MFE | -26.58 |
| Consensus MFE | -10.64 |
| Energy contribution | -11.33 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.40 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729530 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18582360 108 + 20766785 CCAAGAGAAUGAUGUUAGUGUGUGUGUUAGG-UGGGAAACUUGGCUGGGACUGUAGCUAAGUUCAACACCAGGCGGAGAGGCAAUUGUGCAUAGUGGUACAAAGGUUGA ((...............((.(.(.(((..((-((...((((((((((......))))))))))...))))..))).).).))..((((((......)))))).)).... ( -32.40) >DroSim_CAF1 4263 108 + 1 CCAAGAGGAUGAUGUUAGUGUGGAUGUUUGG-UGGGAAACUUGGUUGGGGAUGUAGCUGAGUUCAACACCAGGCGGAGUGGCCAUUGUGCACAGUGGUACAAAGGUUGA (((....(((...)))....))).(((((((-((...((((..((((......))))..))))...)))))))))....((((.((((((......)))))).)))).. ( -33.20) >DroEre_CAF1 7158 83 + 1 CCGAAAG----GUGU-----------AUGGGGUGGAAAACUUGGCUGAGGAUGUAGCAAAGAUCGAAACCAC-----------CGCGUCCACAGUGGUAUAAAAGUAUA (((....----(((.-----------(((.(((((....(((.((((......)))).))).......))))-----------).))).)))..)))............ ( -21.60) >DroYak_CAF1 7306 84 + 1 CCAAAAG----AUAUUAGAGUGGGUGUUGGGGUGGA-AACUUUGCUGAGGAUGUAGCAAGGUUCAAAACC--------------------ACAGUGGUACAAAAAUAUA (((....----.........)))(((..(..((((.-((((((((((......)))))))))).....))--------------------))..)..)))......... ( -19.12) >consensus CCAAAAG____AUGUUAGUGUGGGUGUUGGG_UGGAAAACUUGGCUGAGGAUGUAGCAAAGUUCAAAACCAG___________AUUGUGCACAGUGGUACAAAAGUAGA ..............................((((...((((((((((......))))))))))...))))....................................... (-10.64 = -11.33 + 0.69)
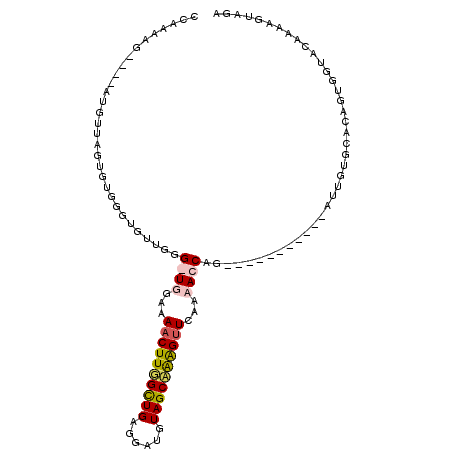


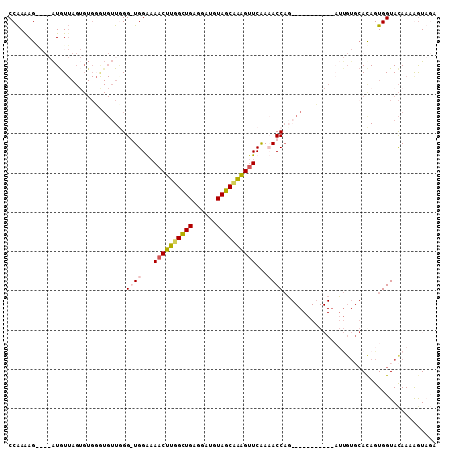
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:12 2006